import numpy as np
import pandas as pd
import os
import json
import requests
import shutil
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.preprocessing import normalize
import plotly.express as px
from _cde_compute_edges_from_nodes import *
pd.set_option('display.max_columns', None)
pd.set_option('display.max_rows', None)
# suppress warnings
import warnings
warnings.filterwarnings("ignore")Distance Analysis: skin-confocal-sorgerlab
Analyze and visualize cell-to-nearest-endothelial-cell distance distributions for the
skin-confocal-sorgerlab dataset.
basepath = "/u/yashjain/hra-cell-distance-analysis/data"
dataset_dir = "skin-confocal-sorgerlab"
data_filedir = os.path.join("data-processed-nodes-with-harmonized-cell-types", dataset_dir)
output_edge_dir = os.path.join("data-processed-edges", dataset_dir)
figures_output_dir = "generated-figures"# Function to load your data
def load_data(path, edges=False):
if edges:
column_names = ['cell_id', 'x1', 'y1', 'z1', 'x2', 'y2', 'z2']
data = pd.read_csv(path, header=None, names=column_names)
else:
data = pd.read_csv(path)
return data# Function to read all files ending with "-nodes.csv" in the `data_filedir` directory into a single DataFrame.
# Another additional column `Dataset` is added to identify the dataset name which comes from the filename before the `-nodes.csv` suffix.
# Additionally, function reads all files ending with "-edges.csv" in the `output_edge_dir` directory into a single DataFrame.
# Three additional columns are added "Dataset", "Anchor Cell Type", and "Anchor Cell Type Level" to identify the dataset name, anchor cell type, and anchor cell type level respectively which come from the filename before the `.csv` suffix.
# The three additional columns are created by splitting the filename on the `-` character, and extracting the relevant parts.
# On splitting, the first part is the dataset name, second part is the anchor cell type level, and third part is the anchor cell type, and last part is the `edges` suffix.
# When reading files, check if the file has the correct format (i.e., ends with `-edges.csv`).
# Additionally, the function merges the edges DataFrame with the nodes DataFrame to get the cell type information for the anchor cells.
# This is done by reading the corresponding nodes file from the `data_filedir` directory for each edges file, and merging it with the edges DataFrame on the `cell_id` column.
# The merged DataFrame contains the edges with additional columns for the cell type information.
# The function returns three DataFrames:
# 1. `merged_nodes`: DataFrame containing all nodes with an additional column `Dataset`.
# 2. `merged_edges`: DataFrame containing all edges with additional columns `Dataset`, `Anchor Cell Type`, and `Anchor Cell Type Level`.
# 3. `merged_nodes_for_all_edges`: DataFrame containing all edges with additional columns `Dataset`, `Anchor Cell Type`, `Anchor Cell Type Level`, and the cell type information for cells.
def read_all_edge_datasets(basepath, data_filedir, output_edge_dir):
all_nodes_files = []
all_edges_files = []
all_nodes_edges_files = []
for file in os.listdir(os.path.join(basepath, output_edge_dir)):
if file.endswith("-edges.csv"):
file_path = os.path.join(basepath, output_edge_dir, file)
dataset_name, anchor_cell_type_level, anchor_cell_type = file.replace("-edges.csv", "").split('-')
edges_df = load_data(file_path, edges=False)
edges_df['Dataset'] = dataset_name
edges_df['Anchor Cell Type'] = anchor_cell_type
edges_df['Anchor Cell Type Level'] = anchor_cell_type_level
edges_df.rename(columns={"distance": "Distance"}, inplace=True) # Rename column "distance" to "Distance".
all_edges_files.append(edges_df)
# Read the corresponding nodes file from data_filedir to get the cell type information
nodes_file_path = os.path.join(basepath, data_filedir, f"{dataset_name}-nodes.csv")
nodes_df = load_data(nodes_file_path)
nodes_df['Dataset'] = dataset_name
all_nodes_files.append(nodes_df)
# Add a new 'cell_id' column to nodes_df
nodes_df['cell_id'] = range(len(nodes_df))
# Set 'cell_id' column as index for nodes_df
nodes_df.set_index('cell_id', inplace=True)
# Merge edges_df with nodes_df to get the cell type information for the anchor cells
edges_nodes_df = pd.merge(edges_df, nodes_df[['Level Three Cell Type', 'Level Two Cell Type', 'Level One Cell Type']], how='left', left_on='cell_id', right_index=True)
all_nodes_edges_files.append(edges_nodes_df)
merged_edges = pd.concat(all_edges_files, ignore_index=True)
merged_nodes = pd.concat(all_nodes_files, ignore_index=True)
merged_nodes_for_all_edges = pd.concat(all_nodes_edges_files, ignore_index=True)
return merged_nodes, merged_edges, merged_nodes_for_all_edgesdef create_directory(directory):
if not os.path.exists(directory):
os.makedirs(directory)
print(f"Directory '{directory}' created successfully.")
else:
print(f"Directory '{directory}' already exists.")Get initial statistics and identify endothelial cell categories for dataset.
df_all_nodes, df_all_edges, df_all_edges_with_cell_types = read_all_edge_datasets(basepath, data_filedir, output_edge_dir)df_all_nodes.head(5)| x | y | z | Original Cell Type | Level Three Cell Type | Level Three CL Label | Level Three CL ID | CL_Match/3 | Level Two Cell Type | Level Two CL Label | Level Two CL ID | CL_Match/2 | Level One Cell Type | Level One CL Label | Level One CL ID | CL_Match/1 | Dataset | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 732.21 | 18.569 | 0.87001 | Unknown | unknown cell | cell:unknown | CL:0000000 | skos:narrowMatch | unknown cell | cell:unknown | CL:0000000 | skos:narrowMatch | unknown cell | cell:unknown | CL:0000000 | skos:narrowMatch | phenotypes_invasive_margin |
| 1 | 733.07 | 22.936 | 4.77910 | Tumor | tumor cell | neoplastic cell | CL:0001063 | skos:exactMatch | abnormal cell | abnormal cell | CL:0001061 | skos:exactMatch | unknown cell | cell:unknown | CL:0000000 | skos:narrowMatch | phenotypes_invasive_margin |
| 2 | 736.84 | 29.739 | 4.43490 | Tumor | tumor cell | neoplastic cell | CL:0001063 | skos:exactMatch | abnormal cell | abnormal cell | CL:0001061 | skos:exactMatch | unknown cell | cell:unknown | CL:0000000 | skos:narrowMatch | phenotypes_invasive_margin |
| 3 | 732.12 | 143.700 | 15.22900 | Tumor | tumor cell | neoplastic cell | CL:0001063 | skos:exactMatch | abnormal cell | abnormal cell | CL:0001061 | skos:exactMatch | unknown cell | cell:unknown | CL:0000000 | skos:narrowMatch | phenotypes_invasive_margin |
| 4 | 714.57 | 544.400 | 1.40060 | Unknown | unknown cell | cell:unknown | CL:0000000 | skos:narrowMatch | unknown cell | cell:unknown | CL:0000000 | skos:narrowMatch | unknown cell | cell:unknown | CL:0000000 | skos:narrowMatch | phenotypes_invasive_margin |
# Print the total number of unique cell types per dataset. Compute separately for each cell type column (Level One Cell Type, Level Two Cell Type, Level Three Cell Type, Original Cell Type).
print("Total number of unique cell types per cell type annnotation level:")
unique_cell_types = {
'Original Cell Type': df_all_nodes['Original Cell Type'].nunique(),
'Level Three Cell Type': df_all_nodes['Level Three Cell Type'].nunique(),
'Level Two Cell Type': df_all_nodes['Level Two Cell Type'].nunique(),
'Level One Cell Type': df_all_nodes['Level One Cell Type'].nunique()
}
for cell_type, count in unique_cell_types.items():
print(f"{cell_type}: {count}")Total number of unique cell types per cell type annnotation level:
Original Cell Type: 15
Level Three Cell Type: 15
Level Two Cell Type: 11
Level One Cell Type: 4# Save the unique cell types containing "endothelial" in name per cell type column (Level One Cell Type, Level Two Cell Type, Level Three Cell Type, Original Cell Type) to a dictionary where the key is the level and the value is a list of unique cell types.
endothelial_cell_types = {
'Original Cell Type': df_all_nodes[df_all_nodes['Original Cell Type'].str.contains("endothelial", case=False, na=False)]['Original Cell Type'].unique().tolist(),
'Level Three Cell Type': df_all_nodes[df_all_nodes['Level Three Cell Type'].str.contains("endothelial", case=False, na=False)]['Level Three Cell Type'].unique().tolist(),
'Level Two Cell Type': df_all_nodes[df_all_nodes['Level Two Cell Type'].str.contains("endothelial", case=False, na=False)]['Level Two Cell Type'].unique().tolist(),
'Level One Cell Type': df_all_nodes[df_all_nodes['Level One Cell Type'].str.contains("endothelial", case=False, na=False)]['Level One Cell Type'].unique().tolist()
}
print("\nEndothelial cell types per cell type annotation level:")
for level, cell_types in endothelial_cell_types.items():
print(f"\n{level}:")
for cell in cell_types:
print(f" - {cell}")
Endothelial cell types per cell type annotation level:
Original Cell Type:
- Endothelial
Level Three Cell Type:
- endothelial cell
Level Two Cell Type:
- endothelial cell
Level One Cell Type:
- endothelial celltype_field_list = ["Level Three Cell Type", "Level Two Cell Type", "Level One Cell Type"] # Skipping Original Cell Type as it is not a hierarchical level.
# Define the anchor cell type (type of endothelial cell) for each level in type_field_list based on available categories in the previous cell. The distance analysis at all three levels will be limited to the specified anchor cell type.
anchor_cell_type_dict = {
'Level Three Cell Type': 'endothelial cell',
'Level Two Cell Type': 'endothelial cell',
'Level One Cell Type': 'endothelial cell'
}Process datasets to add region information to Nodes files.
# Create a dictionary to map skin regions to correct condition.
region_map = {'phenotypes_invasive_margin':'Invasive Margin',
'phenotypes_melanoma_in_situ':'Melanoma In Situ',
}
# Define the standard region sequence for plots
regions = ['Invasive Margin', 'Melanoma In Situ']df_all_nodes.head()| x | y | z | Original Cell Type | Level Three Cell Type | Level Three CL Label | Level Three CL ID | CL_Match/3 | Level Two Cell Type | Level Two CL Label | Level Two CL ID | CL_Match/2 | Level One Cell Type | Level One CL Label | Level One CL ID | CL_Match/1 | Dataset | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 732.21 | 18.569 | 0.87001 | Unknown | unknown cell | cell:unknown | CL:0000000 | skos:narrowMatch | unknown cell | cell:unknown | CL:0000000 | skos:narrowMatch | unknown cell | cell:unknown | CL:0000000 | skos:narrowMatch | phenotypes_invasive_margin |
| 1 | 733.07 | 22.936 | 4.77910 | Tumor | tumor cell | neoplastic cell | CL:0001063 | skos:exactMatch | abnormal cell | abnormal cell | CL:0001061 | skos:exactMatch | unknown cell | cell:unknown | CL:0000000 | skos:narrowMatch | phenotypes_invasive_margin |
| 2 | 736.84 | 29.739 | 4.43490 | Tumor | tumor cell | neoplastic cell | CL:0001063 | skos:exactMatch | abnormal cell | abnormal cell | CL:0001061 | skos:exactMatch | unknown cell | cell:unknown | CL:0000000 | skos:narrowMatch | phenotypes_invasive_margin |
| 3 | 732.12 | 143.700 | 15.22900 | Tumor | tumor cell | neoplastic cell | CL:0001063 | skos:exactMatch | abnormal cell | abnormal cell | CL:0001061 | skos:exactMatch | unknown cell | cell:unknown | CL:0000000 | skos:narrowMatch | phenotypes_invasive_margin |
| 4 | 714.57 | 544.400 | 1.40060 | Unknown | unknown cell | cell:unknown | CL:0000000 | skos:narrowMatch | unknown cell | cell:unknown | CL:0000000 | skos:narrowMatch | unknown cell | cell:unknown | CL:0000000 | skos:narrowMatch | phenotypes_invasive_margin |
# Iterate through the df_all_data dataframe to create new column "Unique Region" based on the "Dataset" column.
# The "Unique Region" column is created by mapping the region names based on the full dataset name.
df_all_nodes['Unique Region'] = df_all_nodes['Dataset'].map(region_map)
# df_all_nodes['Unique Region'] = df_all_nodes['Dataset'].str.split('-').str[1].map(region_map)
# Check if the new columns are created correctly.
df_all_nodes[['Dataset', 'Unique Region']].head(5)| Dataset | Unique Region | |
|---|---|---|
| 0 | phenotypes_invasive_margin | Invasive Margin |
| 1 | phenotypes_invasive_margin | Invasive Margin |
| 2 | phenotypes_invasive_margin | Invasive Margin |
| 3 | phenotypes_invasive_margin | Invasive Margin |
| 4 | phenotypes_invasive_margin | Invasive Margin |
# Print all unique regions in the data.
print("\nUnique Regions in the data:")
print(df_all_nodes['Unique Region'].unique())
# Print the total number of unique regions.
print(f"Total number of unique regions: {df_all_nodes['Unique Region'].nunique()}")
# Print number of unique datasets per unique region.
print("\nNumber of unique datasets per unique region:")
for region in df_all_nodes['Unique Region'].unique():
num_datasets = df_all_nodes[df_all_nodes['Unique Region'] == region]['Dataset'].nunique()
print(f"{region}: {num_datasets}")
Unique Regions in the data:
['Invasive Margin' 'Melanoma In Situ']
Total number of unique regions: 2
Number of unique datasets per unique region:
Invasive Margin: 1
Melanoma In Situ: 1Process datasets to add region information to Edges files.
df_all_edges.head(5)| cell_id | x1 | y1 | z1 | x2 | y2 | z2 | Distance | Dataset | Anchor Cell Type | Anchor Cell Type Level | |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 0 | 732.21 | 18.569 | 0.87001 | 662.23 | 117.79 | 28.219 | 124.458726 | phenotypes_invasive_margin | endothelial cell | Level Three Cell Type |
| 1 | 1 | 733.07 | 22.936 | 4.77910 | 662.23 | 117.79 | 28.219 | 120.685607 | phenotypes_invasive_margin | endothelial cell | Level Three Cell Type |
| 2 | 2 | 736.84 | 29.739 | 4.43490 | 662.23 | 117.79 | 28.219 | 117.835963 | phenotypes_invasive_margin | endothelial cell | Level Three Cell Type |
| 3 | 3 | 732.12 | 143.700 | 15.22900 | 670.45 | 159.88 | 25.529 | 64.583832 | phenotypes_invasive_margin | endothelial cell | Level Three Cell Type |
| 4 | 5 | 748.79 | 80.906 | 12.39500 | 662.23 | 117.79 | 28.219 | 95.412064 | phenotypes_invasive_margin | endothelial cell | Level Three Cell Type |
# Process the edge data to create new columns "Unique Region" based on the "Dataset" column, similar to how it was done for the node data.
df_all_edges['Unique Region'] = df_all_edges['Dataset'].map(region_map)
# Check if the new columns are created correctly.
df_all_edges[['Dataset', 'Unique Region']].head(5)| Dataset | Unique Region | |
|---|---|---|
| 0 | phenotypes_invasive_margin | Invasive Margin |
| 1 | phenotypes_invasive_margin | Invasive Margin |
| 2 | phenotypes_invasive_margin | Invasive Margin |
| 3 | phenotypes_invasive_margin | Invasive Margin |
| 4 | phenotypes_invasive_margin | Invasive Margin |
# Print all unique regions in the data.
print("\nUnique Regions in the data:")
print(df_all_edges['Unique Region'].unique())
# Print the total number of unique regions.
print(f"Total number of unique regions: {df_all_edges['Unique Region'].nunique()}")
# Print number of unique datasets per unique region.
print("\nNumber of unique datasets per unique region:")
for region in df_all_edges['Unique Region'].unique():
num_datasets = df_all_edges[df_all_edges['Unique Region'] == region]['Dataset'].nunique()
print(f"{region}: {num_datasets}")
Unique Regions in the data:
['Invasive Margin' 'Melanoma In Situ']
Total number of unique regions: 2
Number of unique datasets per unique region:
Invasive Margin: 1
Melanoma In Situ: 1df_all_edges_with_cell_types['Unique Region'] = df_all_edges_with_cell_types['Dataset'].map(region_map)
# Check if the new columns are created correctly.
df_all_edges_with_cell_types[['Dataset', 'Unique Region']].head(5)| Dataset | Unique Region | |
|---|---|---|
| 0 | phenotypes_invasive_margin | Invasive Margin |
| 1 | phenotypes_invasive_margin | Invasive Margin |
| 2 | phenotypes_invasive_margin | Invasive Margin |
| 3 | phenotypes_invasive_margin | Invasive Margin |
| 4 | phenotypes_invasive_margin | Invasive Margin |
df_all_nodes.head(1)| x | y | z | Original Cell Type | Level Three Cell Type | Level Three CL Label | Level Three CL ID | CL_Match/3 | Level Two Cell Type | Level Two CL Label | Level Two CL ID | CL_Match/2 | Level One Cell Type | Level One CL Label | Level One CL ID | CL_Match/1 | Dataset | Unique Region | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 732.21 | 18.569 | 0.87001 | Unknown | unknown cell | cell:unknown | CL:0000000 | skos:narrowMatch | unknown cell | cell:unknown | CL:0000000 | skos:narrowMatch | unknown cell | cell:unknown | CL:0000000 | skos:narrowMatch | phenotypes_invasive_margin | Invasive Margin |
df_all_edges.head(1)| cell_id | x1 | y1 | z1 | x2 | y2 | z2 | Distance | Dataset | Anchor Cell Type | Anchor Cell Type Level | Unique Region | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 0 | 732.21 | 18.569 | 0.87001 | 662.23 | 117.79 | 28.219 | 124.458726 | phenotypes_invasive_margin | endothelial cell | Level Three Cell Type | Invasive Margin |
df_all_edges_with_cell_types.head(1)| cell_id | x1 | y1 | z1 | x2 | y2 | z2 | Distance | Dataset | Anchor Cell Type | Anchor Cell Type Level | Level Three Cell Type | Level Two Cell Type | Level One Cell Type | Unique Region | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 0 | 732.21 | 18.569 | 0.87001 | 662.23 | 117.79 | 28.219 | 124.458726 | phenotypes_invasive_margin | endothelial cell | Level Three Cell Type | unknown cell | unknown cell | unknown cell | Invasive Margin |
Node Analysis
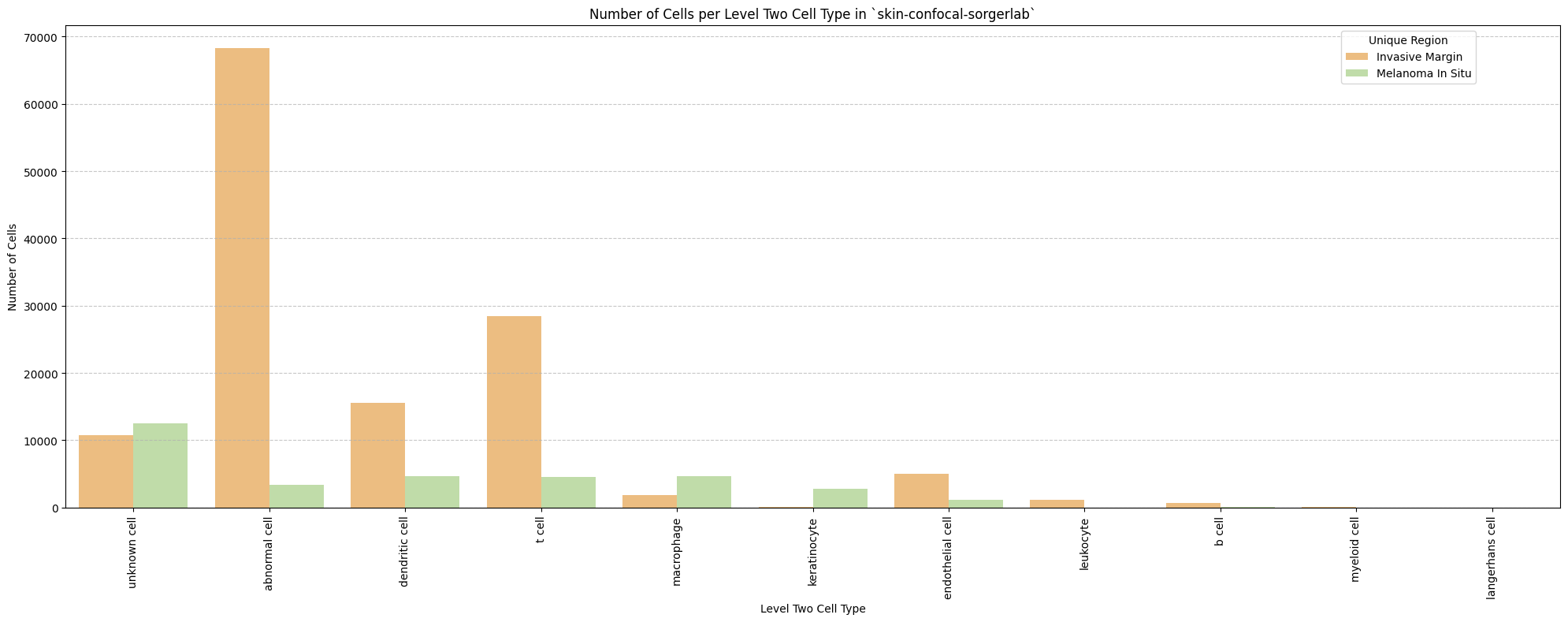
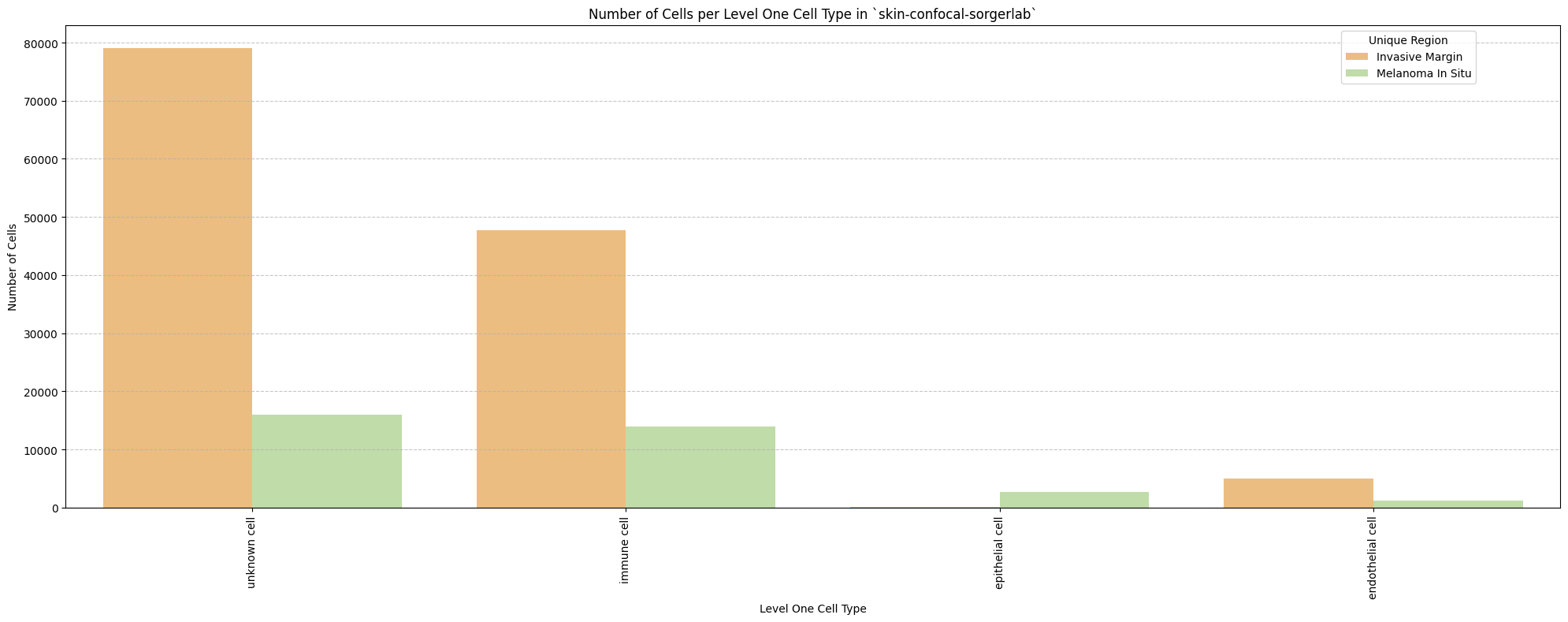
# Plot number of cells per cell type in the same plot. Color by cell type and unique region. Output figure saved in existing `figures_output_dir`.
def plot_cells_per_celltype(df, type_field, output_dir):
plt.figure(figsize=(20, 8))
plt.rcParams["svg.fonttype"] = 'none' # to store text as text, not as path
sns.countplot(data=df, x=type_field, palette='Spectral', hue='Unique Region')
plt.title(f'Number of Cells per {type_field} in `{dataset_dir}`')
plt.xticks(rotation=90)
plt.tight_layout()
plt.savefig(os.path.join(output_dir, f'{dataset_dir}_cells_per_celltype_{type_field}.png'), dpi=300,
bbox_inches='tight',
pad_inches=0.5)
plt.savefig(os.path.join(output_dir, f'{dataset_dir}_cells_per_celltype_{type_field}.svg'), dpi=300,
bbox_inches='tight',
pad_inches=0.5)
plt.legend(title='Unique Region', bbox_to_anchor=(0.85, 1), loc='upper left')
plt.xlabel(type_field)
# For numbers on y-axis, do not use scientific notation.
plt.ticklabel_format(style='plain', axis='y')
# Set y-axis label
plt.ylabel('Number of Cells')
plt.grid(axis='y', linestyle='--', alpha=0.7)
plt.tight_layout()
# Show the plot
plt.show()
plt.close()
for type_field in type_field_list:
plot_cells_per_celltype(df_all_nodes, type_field, os.path.join(basepath, figures_output_dir))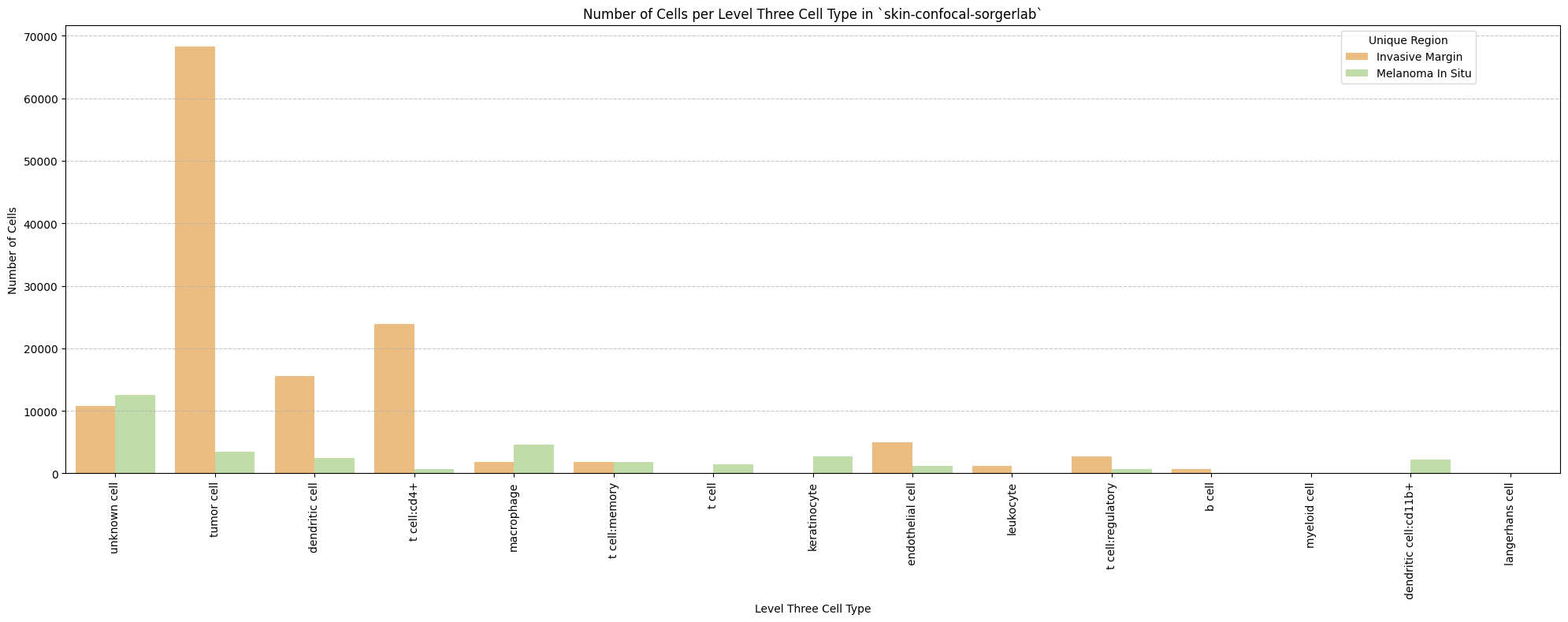
Distance Analysis
# Get mean, median, minimum, maximum distance per unique region per anchor cell type.
df_distance_stats = df_all_edges_with_cell_types.groupby(['Unique Region', 'Anchor Cell Type', 'Anchor Cell Type Level']).agg(
mean_distance=('Distance', 'mean'),
median_distance=('Distance', 'median'),
min_distance=('Distance', 'min'),
max_distance=('Distance', 'max')
).reset_index()
# Print the first few rows of the distance statistics DataFrame.
df_distance_stats| Unique Region | Anchor Cell Type | Anchor Cell Type Level | mean_distance | median_distance | min_distance | max_distance | |
|---|---|---|---|---|---|---|---|
| 0 | Invasive Margin | endothelial cell | Level One Cell Type | 36.952338 | 27.206913 | 1.116692 | 164.060518 |
| 1 | Invasive Margin | endothelial cell | Level Three Cell Type | 36.952338 | 27.206913 | 1.116692 | 164.060518 |
| 2 | Invasive Margin | endothelial cell | Level Two Cell Type | 36.952338 | 27.206913 | 1.116692 | 164.060518 |
| 3 | Melanoma In Situ | endothelial cell | Level One Cell Type | 26.642030 | 20.559508 | 1.857488 | 132.248984 |
| 4 | Melanoma In Situ | endothelial cell | Level Three Cell Type | 26.642030 | 20.559508 | 1.857488 | 132.248984 |
| 5 | Melanoma In Situ | endothelial cell | Level Two Cell Type | 26.642030 | 20.559508 | 1.857488 | 132.248984 |
Level One Cell Type Analysis
# Get mean, median, minimum, maximum distance per cell type in all unique regions.
cell_type_level = 'Level One Cell Type'
df_all_edges_with_cell_type_level = df_all_edges_with_cell_types[(df_all_edges_with_cell_types['Anchor Cell Type Level'] == cell_type_level) & (df_all_edges_with_cell_types['Anchor Cell Type'] == anchor_cell_type_dict[cell_type_level])]
df_distance_stats_cell_type_level = df_all_edges_with_cell_type_level.groupby([cell_type_level, 'Unique Region']).agg(
mean_distance=('Distance', 'mean'),
median_distance=('Distance', 'median'),
min_distance=('Distance', 'min'),
max_distance=('Distance', 'max')
).reset_index()
df_distance_stats_cell_type_level| Level One Cell Type | Unique Region | mean_distance | median_distance | min_distance | max_distance | |
|---|---|---|---|---|---|---|
| 0 | epithelial cell | Invasive Margin | 12.513268 | 9.156887 | 2.954969 | 47.494145 |
| 1 | epithelial cell | Melanoma In Situ | 39.799206 | 36.606082 | 10.294391 | 101.840063 |
| 2 | immune cell | Invasive Margin | 16.909552 | 12.588745 | 1.837015 | 157.657774 |
| 3 | immune cell | Melanoma In Situ | 19.275017 | 14.685411 | 2.029235 | 123.084108 |
| 4 | unknown cell | Invasive Margin | 49.094668 | 41.982022 | 1.116692 | 164.060518 |
| 5 | unknown cell | Melanoma In Situ | 30.822976 | 24.278950 | 1.857488 | 132.248984 |
# Get top five and bottom five cell types with respect to mean distance in each unique region separately.
def get_top_bottom_cell_types_by_mean(df, cell_type_level, unique_region, top_n=5):
# Filter the DataFrame for the specified unique region and cell type level
df_filtered = df[df['Unique Region'] == unique_region]
# Group by the specified cell type level and calculate mean distance
df_grouped = df_filtered.groupby(cell_type_level).agg(mean_distance=('Distance', 'mean')).reset_index()
# Sort by mean distance to get top and bottom cell types
df_sorted = df_grouped.sort_values(by='mean_distance', ascending=False)
# Get top N and bottom N cell types
top_cell_types = df_sorted.head(top_n)
bottom_cell_types = df_sorted.tail(top_n)
return top_cell_types, bottom_cell_types
# Get top and bottom cell types for each unique region in the dataset.
unique_regions = df_all_edges_with_cell_type_level['Unique Region'].unique()
for region in unique_regions:
top_bottom = get_top_bottom_cell_types_by_mean(df_all_edges_with_cell_type_level, cell_type_level, region)
print(f"\nTop 5 cell types in {region}:")
print(top_bottom[0])
print(f"\nBottom 5 cell types in {region}:")
print(top_bottom[1])
Top 5 cell types in Invasive Margin:
Level One Cell Type mean_distance
2 unknown cell 49.094668
1 immune cell 16.909552
0 epithelial cell 12.513268
Bottom 5 cell types in Invasive Margin:
Level One Cell Type mean_distance
2 unknown cell 49.094668
1 immune cell 16.909552
0 epithelial cell 12.513268
Top 5 cell types in Melanoma In Situ:
Level One Cell Type mean_distance
0 epithelial cell 39.799206
2 unknown cell 30.822976
1 immune cell 19.275017
Bottom 5 cell types in Melanoma In Situ:
Level One Cell Type mean_distance
0 epithelial cell 39.799206
2 unknown cell 30.822976
1 immune cell 19.275017# Get top five and bottom five cell types with respect to median distance in each unique region separately.
def get_top_bottom_cell_types_by_median(df, cell_type_level, unique_region, top_n=5):
# Filter the DataFrame for the specified unique region and cell type level
df_filtered = df[df['Unique Region'] == unique_region]
# Group by the specified cell type level and calculate median distance
df_grouped = df_filtered.groupby(cell_type_level).agg(median_distance=('Distance', 'median')).reset_index()
# Sort by median distance to get top and bottom cell types
df_sorted = df_grouped.sort_values(by='median_distance', ascending=False)
# Get top N and bottom N cell types
top_cell_types = df_sorted.head(top_n)
bottom_cell_types = df_sorted.tail(top_n)
return top_cell_types, bottom_cell_types
# Get top and bottom cell types for each unique region in the dataset.
unique_regions = df_all_edges_with_cell_type_level['Unique Region'].unique()
for region in unique_regions:
top_bottom = get_top_bottom_cell_types_by_median(df_all_edges_with_cell_type_level, cell_type_level, region)
print(f"\nTop 5 cell types in {region}:")
print(top_bottom[0])
print(f"\nBottom 5 cell types in {region}:")
print(top_bottom[1])
Top 5 cell types in Invasive Margin:
Level One Cell Type median_distance
2 unknown cell 41.982022
1 immune cell 12.588745
0 epithelial cell 9.156887
Bottom 5 cell types in Invasive Margin:
Level One Cell Type median_distance
2 unknown cell 41.982022
1 immune cell 12.588745
0 epithelial cell 9.156887
Top 5 cell types in Melanoma In Situ:
Level One Cell Type median_distance
0 epithelial cell 36.606082
2 unknown cell 24.278950
1 immune cell 14.685411
Bottom 5 cell types in Melanoma In Situ:
Level One Cell Type median_distance
0 epithelial cell 36.606082
2 unknown cell 24.278950
1 immune cell 14.685411# Calculate regional variability
def calculate_regional_variability(df_all_edges_with_cell_type_level, cell_type_level):
""" Calculate regional variability for distances in the given DataFrame.
"""
regional_variability = df_all_edges_with_cell_type_level.groupby('Unique Region')['Distance'].agg([
('mean', 'mean'),
('std', 'std')
]).round(2)
# Add CV as percentage
regional_variability['CV (%)'] = (regional_variability['std'] / regional_variability['mean'] * 100).round(1)
print("\nRegional Variability Analysis:")
print("Mean: Average distance in each region")
print("Std: Standard deviation of distances")
print("CV: Coefficient of Variation (std/mean * 100%)")
print(regional_variability)
# Calculate variability for each cell type
cell_type_variability = df_all_edges_with_cell_type_level.groupby(cell_type_level)['Distance'].agg([
('mean', 'mean'),
('std', 'std')
]).round(2)
# Add CV as percentage
cell_type_variability['CV (%)'] = (cell_type_variability['std'] / cell_type_variability['mean'] * 100).round(1)
print("\nCell Type Variability Analysis (sorted by CV):")
print(cell_type_variability.sort_values('CV (%)', ascending=False))
calculate_regional_variability(df_all_edges_with_cell_type_level, cell_type_level)
Regional Variability Analysis:
Mean: Average distance in each region
Std: Standard deviation of distances
CV: Coefficient of Variation (std/mean * 100%)
mean std CV (%)
Unique Region
Invasive Margin 36.95 31.48 85.2
Melanoma In Situ 26.64 20.20 75.8
Cell Type Variability Analysis (sorted by CV):
mean std CV (%)
Level One Cell Type
immune cell 17.45 15.16 86.9
unknown cell 46.02 31.84 69.2
epithelial cell 38.94 16.67 42.8# Generate Violin Plot
def plot_violin_cells_per_celltype(df_all_edges_with_cell_type_level, cell_type_level, output_dir, density_norm='area'):
sns.set_style("whitegrid")
sns.set_context("notebook", rc={"grid.linewidth": 2})
plt.figure(figsize=(10, 6))
plt.rcParams["svg.fonttype"] = 'none' # to store text as text, not as path
sns.violinplot(data=df_all_edges_with_cell_type_level, x=cell_type_level, y="Distance", density_norm=density_norm, common_norm=True, cut=0, inner="box", split=False, palette='Spectral', alpha=.9)
sns.set_theme(style="whitegrid")
sns.set_context("paper")
font_size = 10
plt.legend(fontsize=font_size)
plt.title(f'Violin Plot of distances by {cell_type_level} (Density Normalization: {density_norm})', fontsize=font_size)
plt.xlabel(f'{cell_type_level}', fontsize=font_size)
plt.ylabel('Distance (\u03bcm)', fontsize=font_size)
# Increase font size for all text in the figure
plt.xticks(fontsize=font_size)
plt.xticks(rotation=90)
plt.yticks(fontsize=font_size)
plt.tight_layout()
plt.savefig(os.path.join(output_dir, f'{dataset_dir}_violin_cells_per_celltype_{cell_type_level}.png'), dpi=300,
bbox_inches='tight',
pad_inches=0.5)
plt.savefig(os.path.join(output_dir, f'{dataset_dir}_violin_cells_per_celltype_{cell_type_level}.svg'), dpi=300,
bbox_inches='tight',
pad_inches=0.5)
plt.show()
plot_violin_cells_per_celltype(df_all_edges_with_cell_type_level, cell_type_level, os.path.join(basepath, figures_output_dir), density_norm='area')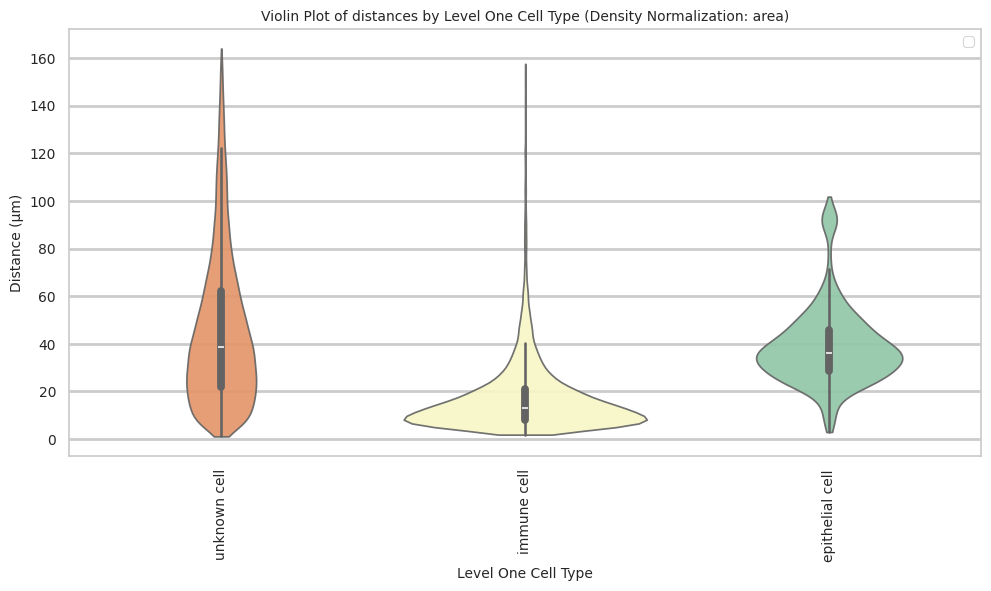
# Boxplots of distribution of distances by cell type and region.
def plot_distance_distribution_boxplots_by_region(df_all_edges_with_cell_type_level, cell_type_level, output_dir):
plt.figure(figsize=(16, 8))
plt.rcParams["svg.fonttype"] = 'none' # to store text as text, not as path
# Create categorical type with only the regions that exist in the data
available_regions = [r for r in regions if r in df_all_edges_with_cell_type_level['Unique Region'].unique()]
df_all_edges_with_cell_type_level['Unique Region'] = pd.Categorical(
df_all_edges_with_cell_type_level['Unique Region'],
categories=available_regions,
ordered=True
)
# Make box plot.
sns.boxplot(data=df_all_edges_with_cell_type_level, x=cell_type_level, y='Distance', hue='Unique Region', showfliers=False, palette='Spectral') # viridis or Spectral palette for better color distinction
font_size = 10
plt.xticks(rotation=90, ha='right', fontsize=font_size)
plt.yticks(fontsize=font_size)
plt.title(f'Distribution of distances by {cell_type_level} and region', fontsize=font_size)
plt.xlabel(f'{cell_type_level}', fontsize=font_size)
plt.ylabel('Distance (\u03bcm)', fontsize=font_size)
plt.legend(bbox_to_anchor=(1, 1), loc='upper left')
plt.tight_layout()
plt.savefig(os.path.join(output_dir, f'{dataset_dir}_distance_distribution_boxplots_by_region_{cell_type_level}.png'), dpi=300,
bbox_inches='tight',
pad_inches=0.5)
plt.savefig(os.path.join(output_dir, f'{dataset_dir}_distance_distribution_boxplots_by_region_{cell_type_level}.svg'), dpi=300,
bbox_inches='tight',
pad_inches=0.5)
plt.show()
plot_distance_distribution_boxplots_by_region(df_all_edges_with_cell_type_level, cell_type_level, os.path.join(basepath, figures_output_dir))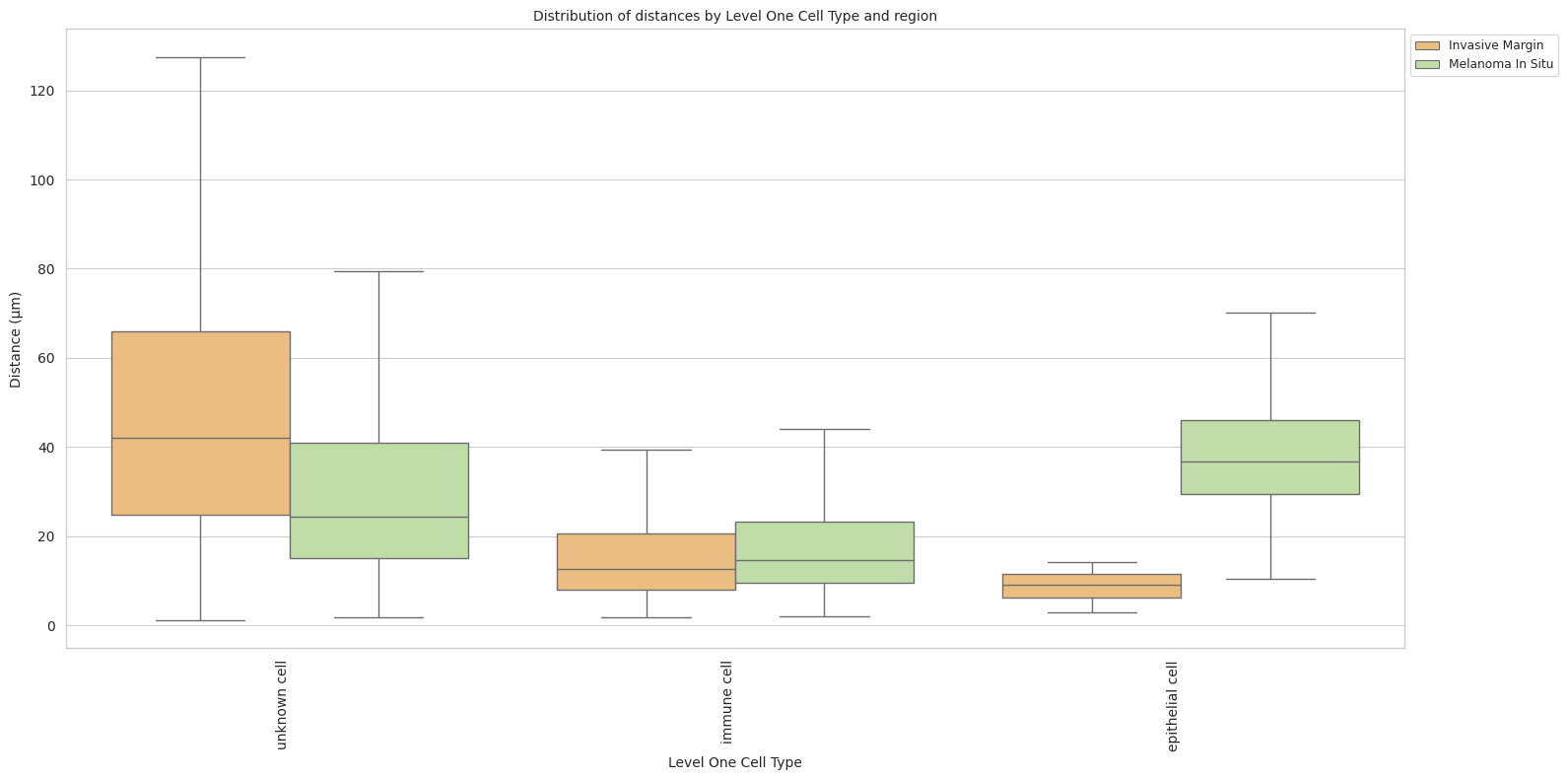
# Boxplots of distribution of distances by cell type and region.
def plot_distance_distribution_heatmap(df_all_edges_with_cell_type_level, cell_type_level, output_dir):
pivot_data = df_all_edges_with_cell_type_level.pivot_table(
values='Distance',
index=cell_type_level,
columns='Unique Region',
aggfunc='median'
)
plt.figure(figsize=(15, 10))
plt.rcParams["svg.fonttype"] = 'none' # to store text as text, not as path
sns.heatmap(pivot_data, annot=True, fmt='.1f', cmap='Spectral')
plt.title(f'Heatmap of median distances by {cell_type_level}', fontsize=12)
font_size = 10
plt.xticks(rotation=90, ha='right', fontsize=font_size)
plt.yticks(fontsize=font_size)
plt.xlabel('Unique Region', fontsize=font_size)
plt.ylabel(f'{cell_type_level}', fontsize=font_size)
plt.tight_layout()
plt.savefig(os.path.join(output_dir, f'{dataset_dir}_distance_distribution_heatmap_{cell_type_level}.png'), dpi=300,
bbox_inches='tight',
pad_inches=0.5)
plt.savefig(os.path.join(output_dir, f'{dataset_dir}_distance_distribution_heatmap_{cell_type_level}.svg'), dpi=300,
bbox_inches='tight',
pad_inches=0.5)
plt.show()
plot_distance_distribution_heatmap(df_all_edges_with_cell_type_level, cell_type_level, os.path.join(basepath, figures_output_dir))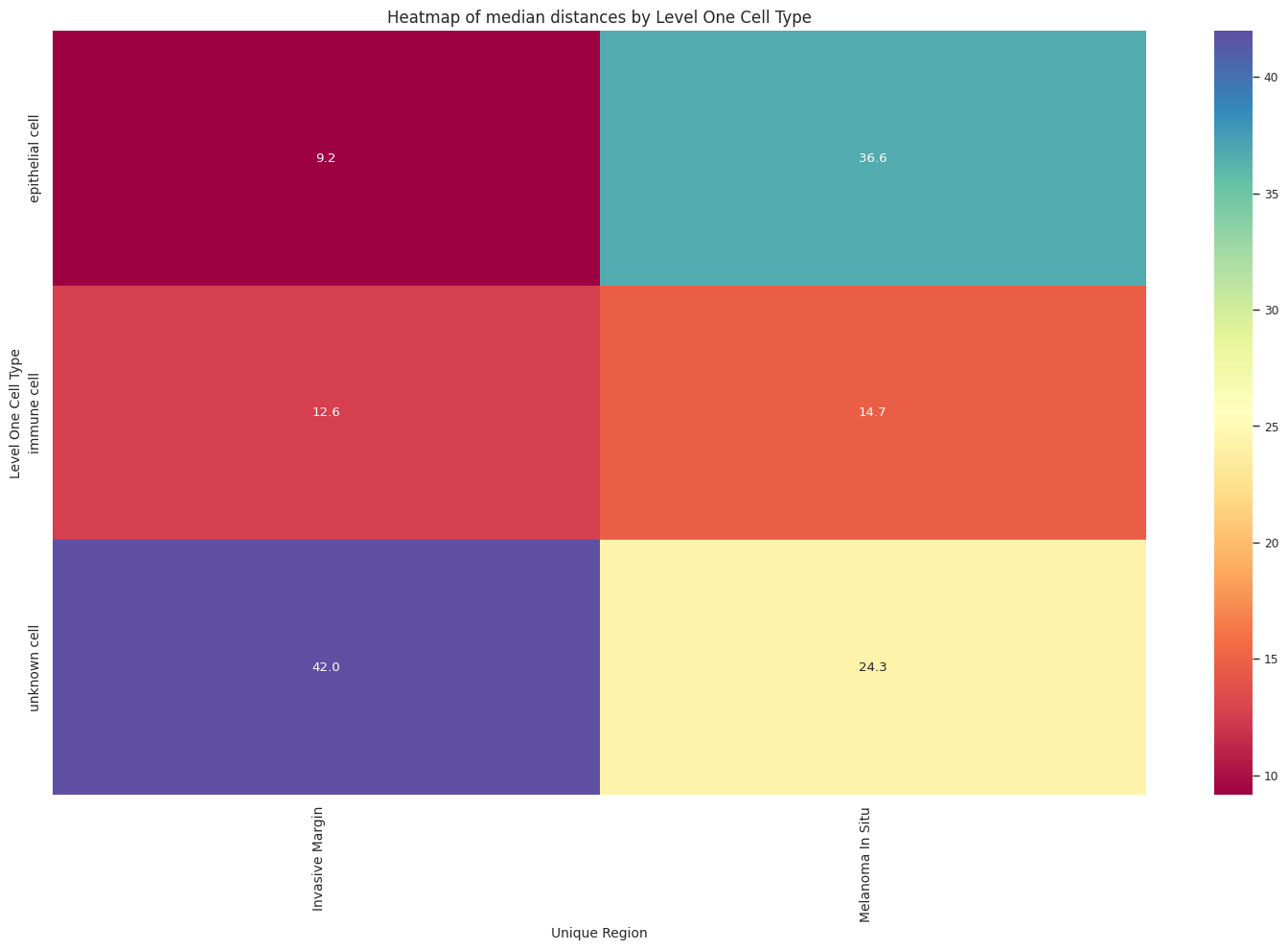
# Generate Violin Plot per unique region in both small intestine and large intestine. Create for all 8 regions as 8 subplots.
def plot_violin_plots_all_regions(df_all_edges_with_cell_type_level, cell_type_level, output_dir, density_norm="area"):
sns.set_style("whitegrid")
sns.set_context("notebook", rc={"grid.linewidth": 1})
plt.rcParams["svg.fonttype"] = 'none' # to store text as text, not as path
font_size = 10
fig, axs = plt.subplots(2, 1, figsize=(10, 15)) # Adjusted figsize for horizontal layout
fig.suptitle(f'Distance distribution per {cell_type_level} in `{dataset_dir}` (density normalization = {density_norm})', fontsize=font_size, y=1)
# Keep the sequence of Cell Types consistent across plots.
cell_types = sorted(df_all_edges_with_cell_type_level[cell_type_level].unique())
# Create a color palette based on the number of unique classes
color_palette = sns.color_palette("Spectral", n_colors=len(cell_types))
# Create a dictionary mapping class to color
class_color_dict = dict(zip(cell_types, color_palette))
for i, region in enumerate(regions):
data_reg = df_all_edges_with_cell_type_level[df_all_edges_with_cell_type_level['Unique Region'] == region]
sns.violinplot(data=data_reg, x=cell_type_level, y="Distance", density_norm=density_norm, common_norm=True, cut=0, inner="box", split=False, palette=class_color_dict, alpha=.9, ax=axs[i], hue=cell_type_level, legend=False, order=cell_types, fill=True)
axs[i].set_title(region, fontsize=font_size)
axs[i].set_xlabel('', fontsize=font_size)
axs[i].set_ylabel('Distance (\u03bcm)', fontsize=font_size)
# axs[i].tick_params(axis='x', labelrotation=90, labelsize=font_size)
# only show xtick labels for the last subplot
if i < len(regions) - 1:
axs[i].set_xticklabels([])
else:
axs[i].set_xticklabels(cell_types, fontsize=font_size, rotation=90, ha='right')
# axs[i].set_ylim(0, data_reg['Distance'].max() * 1.1) # Set y-limits to be consistent across all plots
axs[i].tick_params(axis='both', labelsize=font_size)
# Use fig.text for precise label positioning
fig.figure.text(0.5, -0.02, f'{cell_type_level}', ha='center', va='bottom', fontsize=font_size)
plt.tight_layout()
plt.savefig(os.path.join(output_dir, f'{dataset_dir}_violin_plots_all_regions_{cell_type_level}.png'), dpi=300,
bbox_inches='tight',
pad_inches=0.5)
plt.savefig(os.path.join(output_dir, f'{dataset_dir}_violin_plots_all_regions_{cell_type_level}.svg'), dpi=300,
bbox_inches='tight',
pad_inches=0.5)
plt.show()
plot_violin_plots_all_regions(df_all_edges_with_cell_type_level, cell_type_level, os.path.join(basepath, figures_output_dir), density_norm="count") # density_norm="count" or "area" can be used based on preference.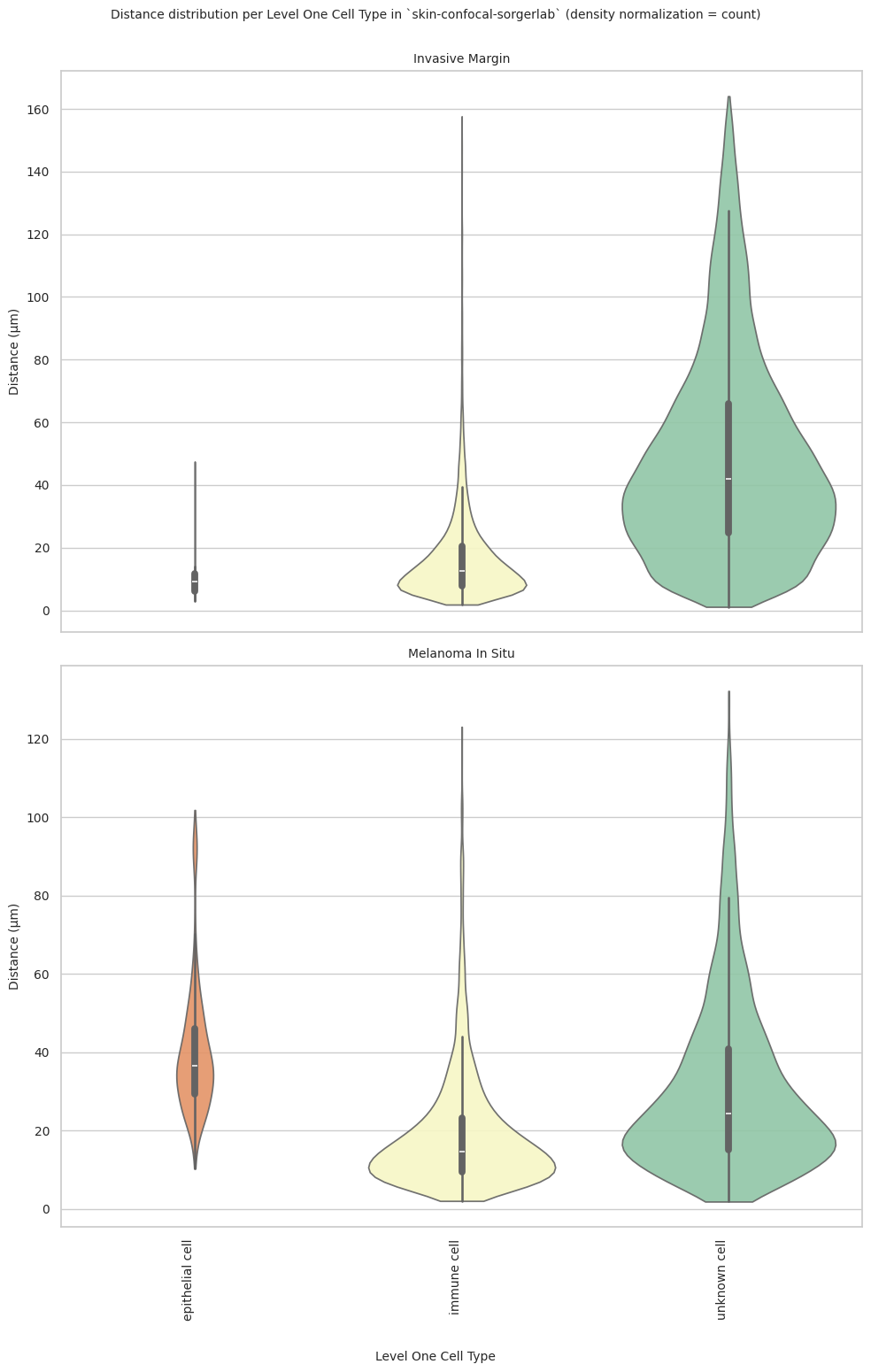
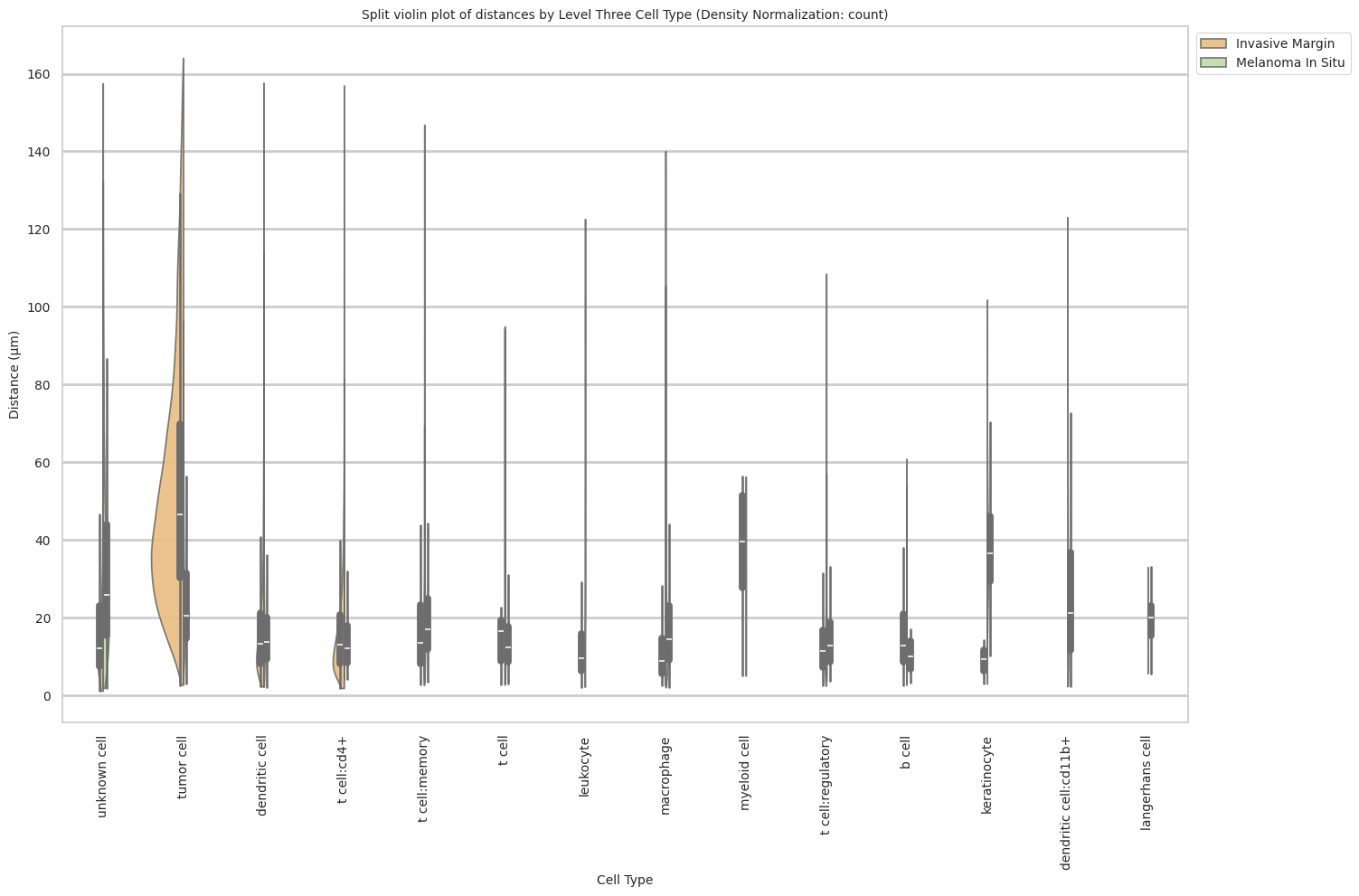
# Generate Split Violin Plot
def plot_violin_cells_per_celltype_split_by_condition(df_all_edges_with_cell_type_level, cell_type_level, output_dir, density_norm="area"):
sns.set_style("whitegrid")
sns.set_context("notebook", rc={"grid.linewidth": 2})
plt.figure(figsize=(15, 10))
plt.rcParams["svg.fonttype"] = 'none' # to store text as text, not as path
sns.violinplot(data=df_all_edges_with_cell_type_level, x=cell_type_level, y="Distance", hue="Unique Region", density_norm=density_norm, common_norm=True, cut=0, inner="box", split=True, palette='Spectral', alpha=.9, hue_order=regions)
sns.set_theme(style="whitegrid")
sns.set_context("paper")
font_size = 10
plt.legend(fontsize=font_size, loc='upper right', bbox_to_anchor=(1.15, 1))
plt.title(f'Split violin plot of distances by {cell_type_level} (Density Normalization: {density_norm})', fontsize=font_size)
plt.xlabel('Cell Type', fontsize=font_size)
plt.ylabel('Distance (\u03bcm)', fontsize=font_size)
# Increase font size for all text in the figure
plt.xticks(fontsize=font_size)
plt.xticks(rotation=90)
plt.yticks(fontsize=font_size)
plt.tight_layout()
plt.savefig(os.path.join(output_dir, f'{dataset_dir}_plot_violin_cells_per_celltype_split_by_condition_{cell_type_level}.png'), dpi=300,
bbox_inches='tight',
pad_inches=0.5)
plt.savefig(os.path.join(output_dir, f'{dataset_dir}_plot_violin_cells_per_celltype_split_by_condition_{cell_type_level}.svg'), dpi=300,
bbox_inches='tight',
pad_inches=0.5)
plt.show()
plot_violin_cells_per_celltype_split_by_condition(df_all_edges_with_cell_type_level, cell_type_level, os.path.join(basepath, figures_output_dir), density_norm="count") # density_norm="count" or "area" can be used based on preference.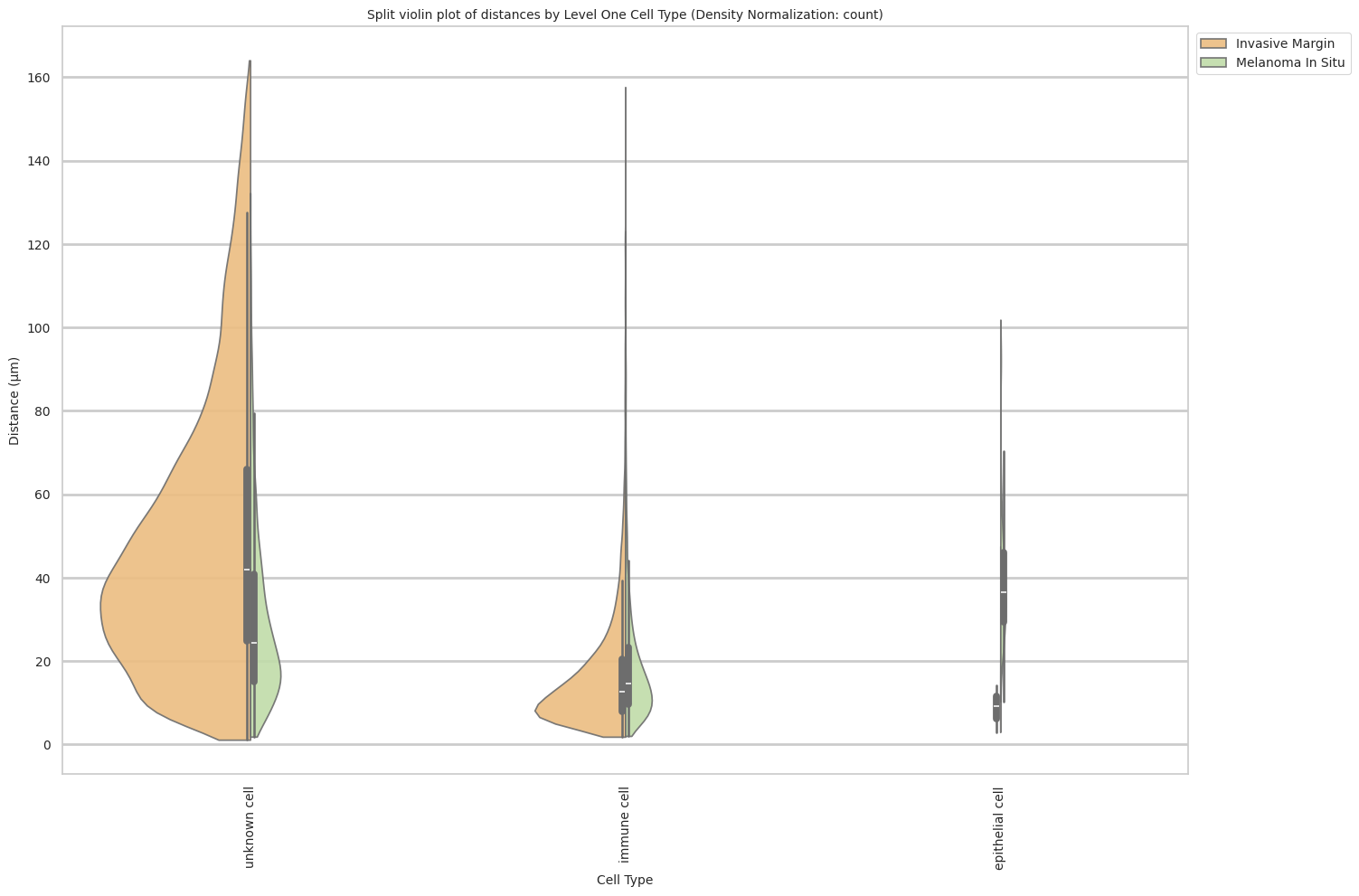
Level Two Cell Type Analysis
# Get mean, median, minimum, maximum distance per cell type in all unique regions.
cell_type_level = 'Level Two Cell Type'
df_all_edges_with_cell_type_level = df_all_edges_with_cell_types[(df_all_edges_with_cell_types['Anchor Cell Type Level'] == cell_type_level) & (df_all_edges_with_cell_types['Anchor Cell Type'] == anchor_cell_type_dict[cell_type_level])]
df_distance_stats_cell_type_level = df_all_edges_with_cell_type_level.groupby([cell_type_level, 'Unique Region']).agg(
mean_distance=('Distance', 'mean'),
median_distance=('Distance', 'median'),
min_distance=('Distance', 'min'),
max_distance=('Distance', 'max')
).reset_index()
df_distance_stats_cell_type_level| Level Two Cell Type | Unique Region | mean_distance | median_distance | min_distance | max_distance | |
|---|---|---|---|---|---|---|
| 0 | abnormal cell | Invasive Margin | 53.754723 | 46.413112 | 2.627413 | 164.060518 |
| 1 | abnormal cell | Melanoma In Situ | 24.330461 | 20.336106 | 3.055009 | 96.398198 |
| 2 | b cell | Invasive Margin | 16.133313 | 12.748624 | 2.619609 | 60.873911 |
| 3 | b cell | Melanoma In Situ | 13.188554 | 9.952653 | 3.154159 | 54.089116 |
| 4 | dendritic cell | Invasive Margin | 18.041947 | 13.133429 | 2.264001 | 157.657774 |
| 5 | dendritic cell | Melanoma In Situ | 21.917961 | 16.043931 | 2.149989 | 123.084108 |
| 6 | keratinocyte | Invasive Margin | 12.513268 | 9.156887 | 2.954969 | 47.494145 |
| 7 | keratinocyte | Melanoma In Situ | 39.799206 | 36.606082 | 10.294391 | 101.840063 |
| 8 | langerhans cell | Melanoma In Situ | 19.308598 | 19.872430 | 5.592065 | 32.988076 |
| 9 | leukocyte | Invasive Margin | 14.150628 | 9.487645 | 2.133846 | 122.610377 |
| 10 | macrophage | Invasive Margin | 12.816781 | 8.688118 | 2.553700 | 140.116350 |
| 11 | macrophage | Melanoma In Situ | 19.304383 | 14.321913 | 2.029235 | 105.486349 |
| 12 | myeloid cell | Invasive Margin | 36.488780 | 39.542376 | 4.989397 | 56.290766 |
| 13 | t cell | Invasive Margin | 16.649191 | 12.753359 | 1.837015 | 156.967478 |
| 14 | t cell | Melanoma In Situ | 16.647669 | 14.136532 | 2.877093 | 94.944415 |
| 15 | unknown cell | Invasive Margin | 19.548098 | 12.104844 | 1.116692 | 157.564532 |
| 16 | unknown cell | Melanoma In Situ | 32.603833 | 25.717688 | 1.857488 | 132.248984 |
# Get top and bottom cell types for each unique region in the dataset.
unique_regions = df_all_edges_with_cell_type_level['Unique Region'].unique()
for region in unique_regions:
top_bottom = get_top_bottom_cell_types_by_mean(df_all_edges_with_cell_type_level, cell_type_level, region)
print(f"\nTop 5 cell types in {region}:")
print(top_bottom[0])
print(f"\nBottom 5 cell types in {region}:")
print(top_bottom[1])
Top 5 cell types in Invasive Margin:
Level Two Cell Type mean_distance
0 abnormal cell 53.754723
6 myeloid cell 36.488780
8 unknown cell 19.548098
2 dendritic cell 18.041947
7 t cell 16.649191
Bottom 5 cell types in Invasive Margin:
Level Two Cell Type mean_distance
7 t cell 16.649191
1 b cell 16.133313
4 leukocyte 14.150628
5 macrophage 12.816781
3 keratinocyte 12.513268
Top 5 cell types in Melanoma In Situ:
Level Two Cell Type mean_distance
3 keratinocyte 39.799206
7 unknown cell 32.603833
0 abnormal cell 24.330461
2 dendritic cell 21.917961
4 langerhans cell 19.308598
Bottom 5 cell types in Melanoma In Situ:
Level Two Cell Type mean_distance
2 dendritic cell 21.917961
4 langerhans cell 19.308598
5 macrophage 19.304383
6 t cell 16.647669
1 b cell 13.188554# Get top and bottom cell types for each unique region in the dataset.
unique_regions = df_all_edges_with_cell_type_level['Unique Region'].unique()
for region in unique_regions:
top_bottom = get_top_bottom_cell_types_by_median(df_all_edges_with_cell_type_level, cell_type_level, region)
print(f"\nTop 5 cell types in {region}:")
print(top_bottom[0])
print(f"\nBottom 5 cell types in {region}:")
print(top_bottom[1])
Top 5 cell types in Invasive Margin:
Level Two Cell Type median_distance
0 abnormal cell 46.413112
6 myeloid cell 39.542376
2 dendritic cell 13.133429
7 t cell 12.753359
1 b cell 12.748624
Bottom 5 cell types in Invasive Margin:
Level Two Cell Type median_distance
1 b cell 12.748624
8 unknown cell 12.104844
4 leukocyte 9.487645
3 keratinocyte 9.156887
5 macrophage 8.688118
Top 5 cell types in Melanoma In Situ:
Level Two Cell Type median_distance
3 keratinocyte 36.606082
7 unknown cell 25.717688
0 abnormal cell 20.336106
4 langerhans cell 19.872430
2 dendritic cell 16.043931
Bottom 5 cell types in Melanoma In Situ:
Level Two Cell Type median_distance
4 langerhans cell 19.872430
2 dendritic cell 16.043931
5 macrophage 14.321913
6 t cell 14.136532
1 b cell 9.952653calculate_regional_variability(df_all_edges_with_cell_type_level, cell_type_level)
Regional Variability Analysis:
Mean: Average distance in each region
Std: Standard deviation of distances
CV: Coefficient of Variation (std/mean * 100%)
mean std CV (%)
Unique Region
Invasive Margin 36.95 31.48 85.2
Melanoma In Situ 26.64 20.20 75.8
Cell Type Variability Analysis (sorted by CV):
mean std CV (%)
Level Two Cell Type
leukocyte 14.15 14.65 103.5
dendritic cell 18.93 17.41 92.0
macrophage 17.47 15.38 88.0
unknown cell 26.57 23.34 87.8
t cell 16.65 13.55 81.4
b cell 15.89 11.25 70.8
abnormal cell 52.34 31.67 60.5
myeloid cell 36.49 16.89 46.3
keratinocyte 38.94 16.67 42.8
langerhans cell 19.31 6.59 34.1plot_violin_cells_per_celltype(df_all_edges_with_cell_type_level, cell_type_level, os.path.join(basepath, figures_output_dir), density_norm='area')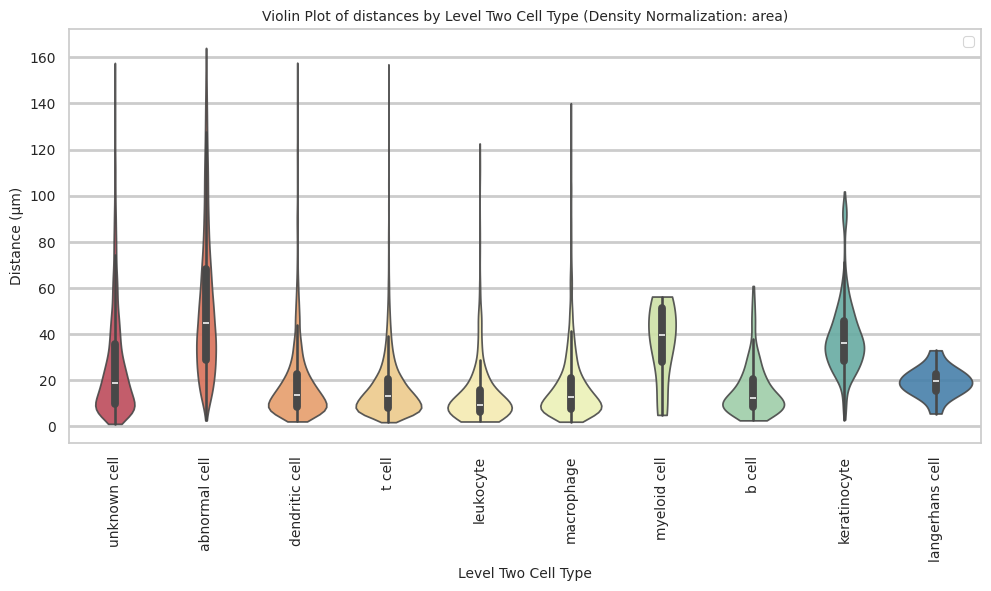
plot_distance_distribution_boxplots_by_region(df_all_edges_with_cell_type_level, cell_type_level, os.path.join(basepath, figures_output_dir))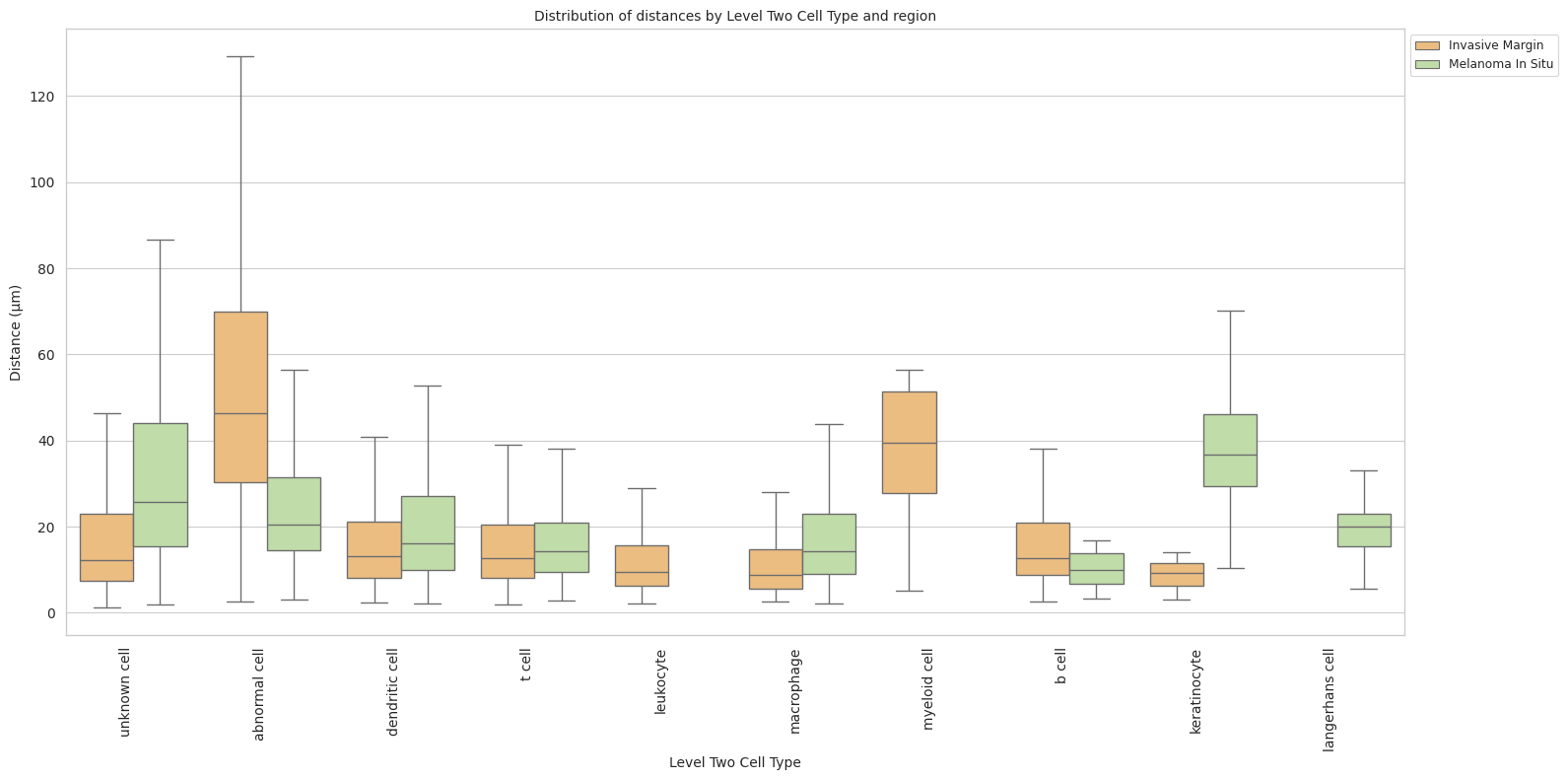
plot_distance_distribution_heatmap(df_all_edges_with_cell_type_level, cell_type_level, os.path.join(basepath, figures_output_dir))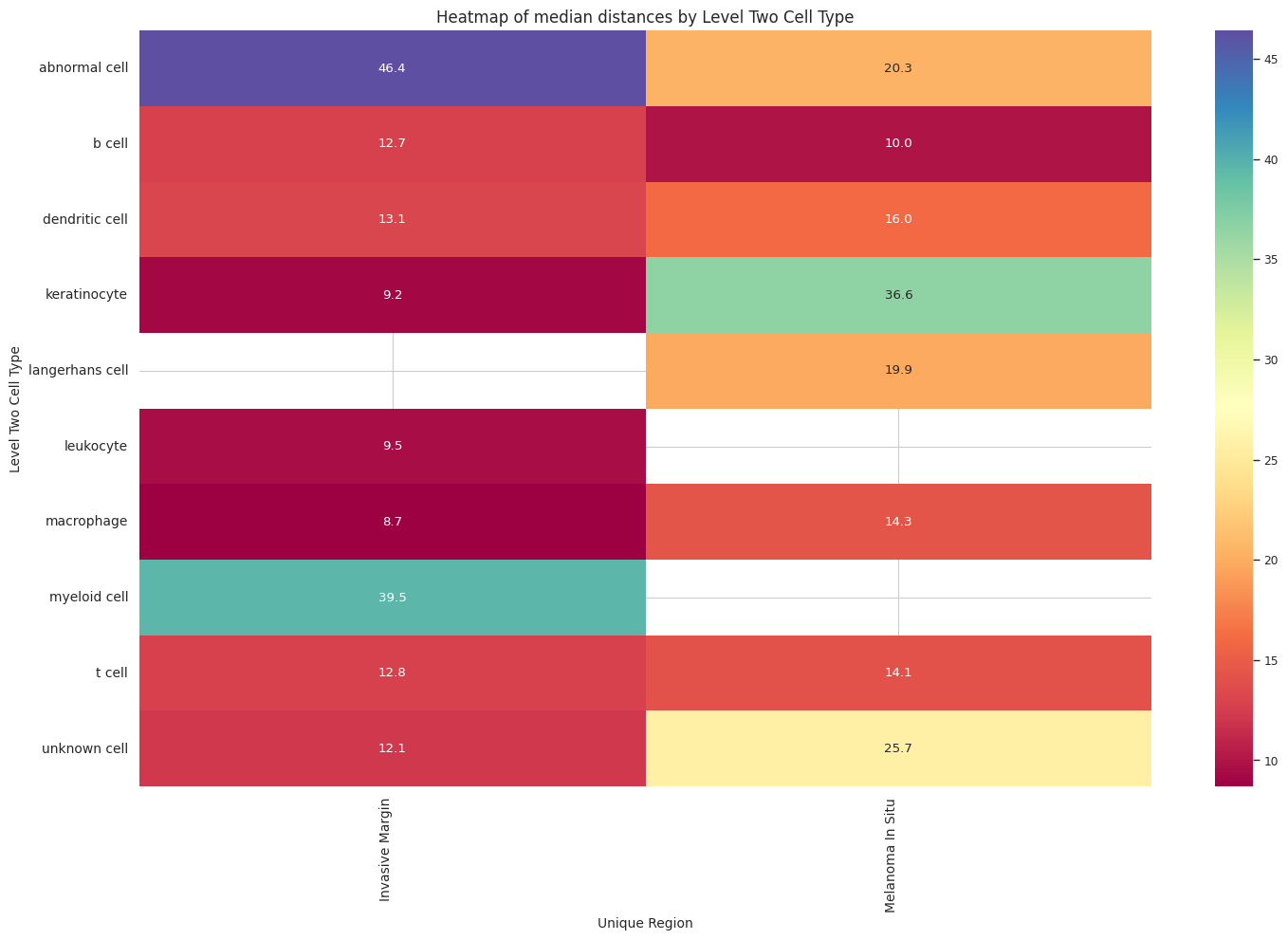
plot_violin_plots_all_regions(df_all_edges_with_cell_type_level, cell_type_level, os.path.join(basepath, figures_output_dir), density_norm="count") # Or, density_norm="count" or "area" based on preference.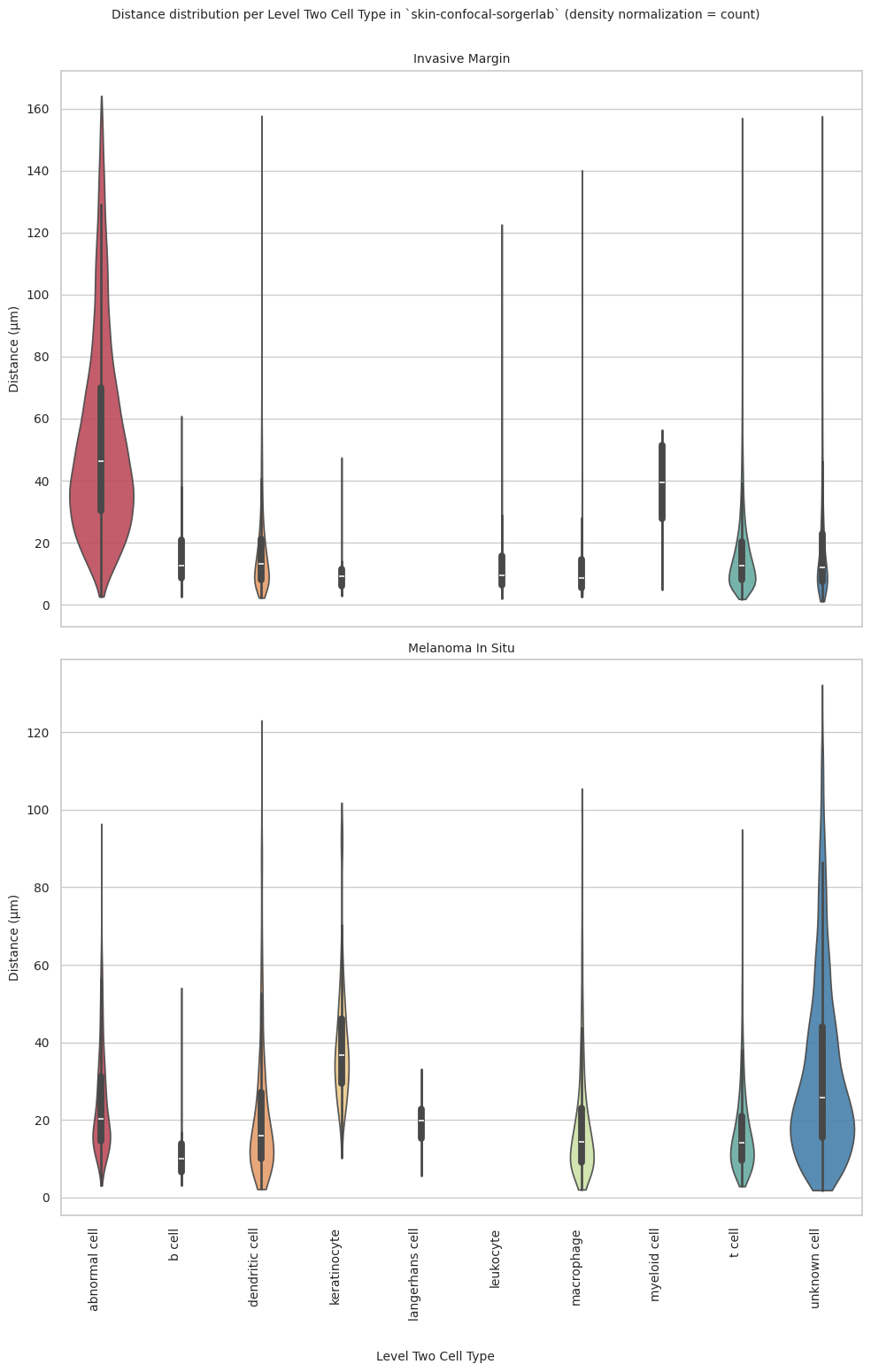
plot_violin_cells_per_celltype_split_by_condition(df_all_edges_with_cell_type_level, cell_type_level, os.path.join(basepath, figures_output_dir), density_norm="count") # density_norm="count" or "area" can be used based on preference.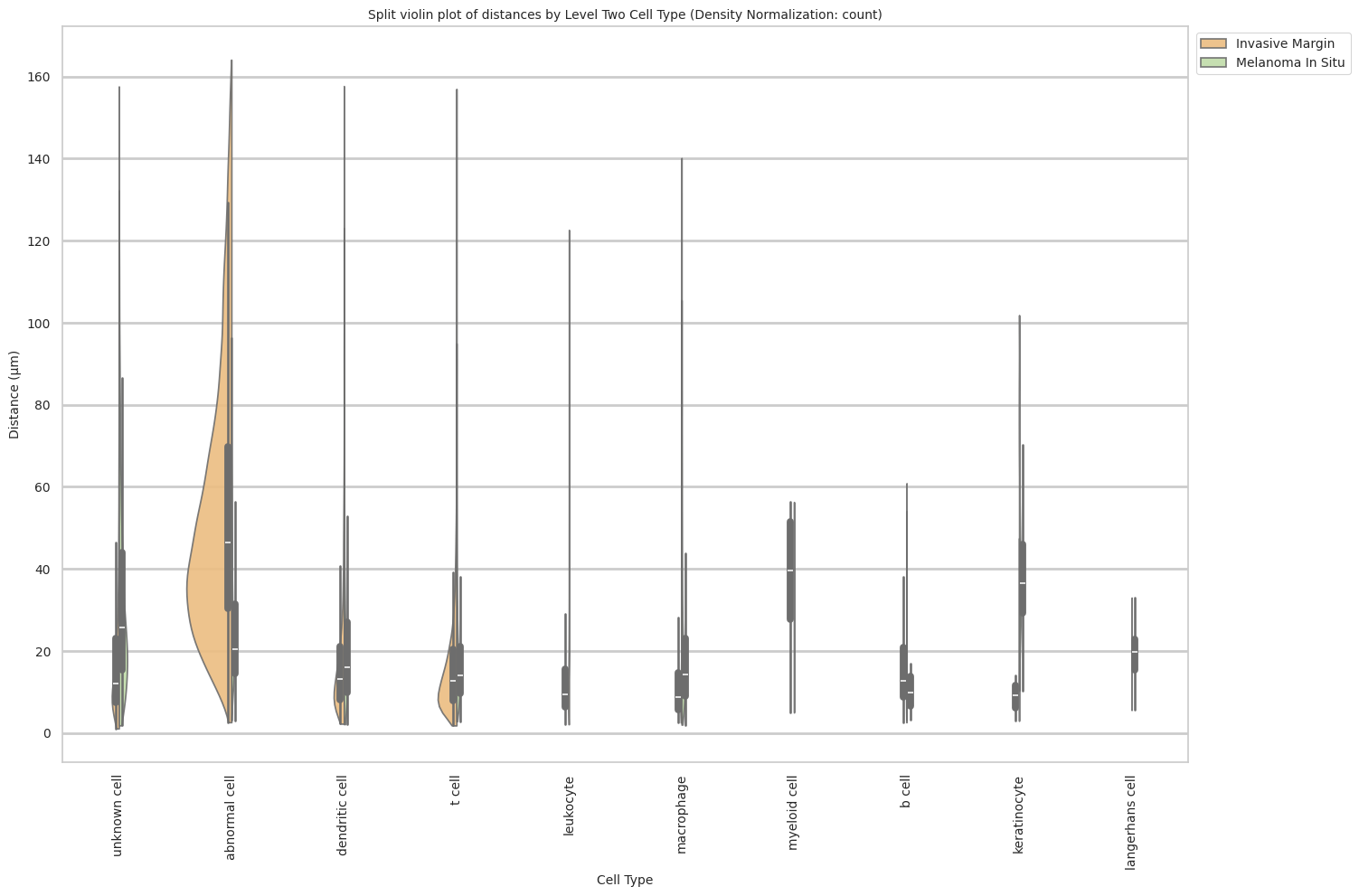
Level Three Cell Type Analysis
# Get mean, median, minimum, maximum distance per cell type in all unique regions.
cell_type_level = 'Level Three Cell Type'
df_all_edges_with_cell_type_level = df_all_edges_with_cell_types[(df_all_edges_with_cell_types['Anchor Cell Type Level'] == cell_type_level) & (df_all_edges_with_cell_types['Anchor Cell Type'] == anchor_cell_type_dict[cell_type_level])]
df_distance_stats_cell_type_level = df_all_edges_with_cell_type_level.groupby([cell_type_level, 'Unique Region']).agg(
mean_distance=('Distance', 'mean'),
median_distance=('Distance', 'median'),
min_distance=('Distance', 'min'),
max_distance=('Distance', 'max')
).reset_index()
df_distance_stats_cell_type_level| Level Three Cell Type | Unique Region | mean_distance | median_distance | min_distance | max_distance | |
|---|---|---|---|---|---|---|
| 0 | b cell | Invasive Margin | 16.133313 | 12.748624 | 2.619609 | 60.873911 |
| 1 | b cell | Melanoma In Situ | 13.188554 | 9.952653 | 3.154159 | 54.089116 |
| 2 | dendritic cell | Invasive Margin | 18.041947 | 13.133429 | 2.264001 | 157.657774 |
| 3 | dendritic cell | Melanoma In Situ | 16.444097 | 13.720679 | 2.149989 | 113.986184 |
| 4 | dendritic cell:cd11b+ | Melanoma In Situ | 27.756263 | 21.201260 | 2.307007 | 123.084108 |
| 5 | keratinocyte | Invasive Margin | 12.513268 | 9.156887 | 2.954969 | 47.494145 |
| 6 | keratinocyte | Melanoma In Situ | 39.799206 | 36.606082 | 10.294391 | 101.840063 |
| 7 | langerhans cell | Melanoma In Situ | 19.308598 | 19.872430 | 5.592065 | 32.988076 |
| 8 | leukocyte | Invasive Margin | 14.150628 | 9.487645 | 2.133846 | 122.610377 |
| 9 | macrophage | Invasive Margin | 12.816781 | 8.688118 | 2.553700 | 140.116350 |
| 10 | macrophage | Melanoma In Situ | 19.304383 | 14.321913 | 2.029235 | 105.486349 |
| 11 | myeloid cell | Invasive Margin | 36.488780 | 39.542376 | 4.989397 | 56.290766 |
| 12 | t cell | Invasive Margin | 18.817402 | 16.529503 | 2.679228 | 94.281492 |
| 13 | t cell | Melanoma In Situ | 14.458865 | 12.312060 | 2.877093 | 94.944415 |
| 14 | t cell:cd4+ | Invasive Margin | 16.783732 | 12.927457 | 1.837015 | 156.967478 |
| 15 | t cell:cd4+ | Melanoma In Situ | 15.262424 | 12.078852 | 4.083577 | 81.476293 |
| 16 | t cell:memory | Invasive Margin | 18.882762 | 13.510906 | 2.649535 | 146.884635 |
| 17 | t cell:memory | Melanoma In Situ | 19.229108 | 17.045629 | 3.338121 | 69.390221 |
| 18 | t cell:regulatory | Invasive Margin | 13.805652 | 11.268087 | 2.410394 | 108.511577 |
| 19 | t cell:regulatory | Melanoma In Situ | 15.844224 | 12.851213 | 3.645520 | 56.987161 |
| 20 | tumor cell | Invasive Margin | 53.754723 | 46.413112 | 2.627413 | 164.060518 |
| 21 | tumor cell | Melanoma In Situ | 24.330461 | 20.336106 | 3.055009 | 96.398198 |
| 22 | unknown cell | Invasive Margin | 19.548098 | 12.104844 | 1.116692 | 157.564532 |
| 23 | unknown cell | Melanoma In Situ | 32.603833 | 25.717688 | 1.857488 | 132.248984 |
# Get top and bottom cell types for each unique region in the dataset.
unique_regions = df_all_edges_with_cell_type_level['Unique Region'].unique()
for region in unique_regions:
top_bottom = get_top_bottom_cell_types_by_mean(df_all_edges_with_cell_type_level, cell_type_level, region)
print(f"\nTop 5 cell types in {region}:")
print(top_bottom[0])
print(f"\nBottom 5 cell types in {region}:")
print(top_bottom[1])
Top 5 cell types in Invasive Margin:
Level Three Cell Type mean_distance
10 tumor cell 53.754723
5 myeloid cell 36.488780
11 unknown cell 19.548098
8 t cell:memory 18.882762
6 t cell 18.817402
Bottom 5 cell types in Invasive Margin:
Level Three Cell Type mean_distance
0 b cell 16.133313
3 leukocyte 14.150628
9 t cell:regulatory 13.805652
4 macrophage 12.816781
2 keratinocyte 12.513268
Top 5 cell types in Melanoma In Situ:
Level Three Cell Type mean_distance
3 keratinocyte 39.799206
11 unknown cell 32.603833
2 dendritic cell:cd11b+ 27.756263
10 tumor cell 24.330461
4 langerhans cell 19.308598
Bottom 5 cell types in Melanoma In Situ:
Level Three Cell Type mean_distance
1 dendritic cell 16.444097
9 t cell:regulatory 15.844224
7 t cell:cd4+ 15.262424
6 t cell 14.458865
0 b cell 13.188554# Get top and bottom cell types for each unique region in the dataset.
unique_regions = df_all_edges_with_cell_type_level['Unique Region'].unique()
for region in unique_regions:
top_bottom = get_top_bottom_cell_types_by_median(df_all_edges_with_cell_type_level, cell_type_level, region)
print(f"\nTop 5 cell types in {region}:")
print(top_bottom[0])
print(f"\nBottom 5 cell types in {region}:")
print(top_bottom[1])
Top 5 cell types in Invasive Margin:
Level Three Cell Type median_distance
10 tumor cell 46.413112
5 myeloid cell 39.542376
6 t cell 16.529503
8 t cell:memory 13.510906
1 dendritic cell 13.133429
Bottom 5 cell types in Invasive Margin:
Level Three Cell Type median_distance
11 unknown cell 12.104844
9 t cell:regulatory 11.268087
3 leukocyte 9.487645
2 keratinocyte 9.156887
4 macrophage 8.688118
Top 5 cell types in Melanoma In Situ:
Level Three Cell Type median_distance
3 keratinocyte 36.606082
11 unknown cell 25.717688
2 dendritic cell:cd11b+ 21.201260
10 tumor cell 20.336106
4 langerhans cell 19.872430
Bottom 5 cell types in Melanoma In Situ:
Level Three Cell Type median_distance
1 dendritic cell 13.720679
9 t cell:regulatory 12.851213
6 t cell 12.312060
7 t cell:cd4+ 12.078852
0 b cell 9.952653calculate_regional_variability(df_all_edges_with_cell_type_level, cell_type_level)
Regional Variability Analysis:
Mean: Average distance in each region
Std: Standard deviation of distances
CV: Coefficient of Variation (std/mean * 100%)
mean std CV (%)
Unique Region
Invasive Margin 36.95 31.48 85.2
Melanoma In Situ 26.64 20.20 75.8
Cell Type Variability Analysis (sorted by CV):
mean std CV (%)
Level Three Cell Type
leukocyte 14.15 14.65 103.5
dendritic cell 17.83 16.38 91.9
macrophage 17.47 15.38 88.0
unknown cell 26.57 23.34 87.8
t cell:cd4+ 16.74 13.80 82.4
dendritic cell:cd11b+ 27.76 22.23 80.1
t cell:memory 19.05 14.91 78.3
t cell:regulatory 14.21 10.79 75.9
b cell 15.89 11.25 70.8
t cell 14.65 9.80 66.9
tumor cell 52.34 31.67 60.5
myeloid cell 36.49 16.89 46.3
keratinocyte 38.94 16.67 42.8
langerhans cell 19.31 6.59 34.1plot_violin_cells_per_celltype(df_all_edges_with_cell_type_level, cell_type_level, os.path.join(basepath, figures_output_dir), density_norm='area')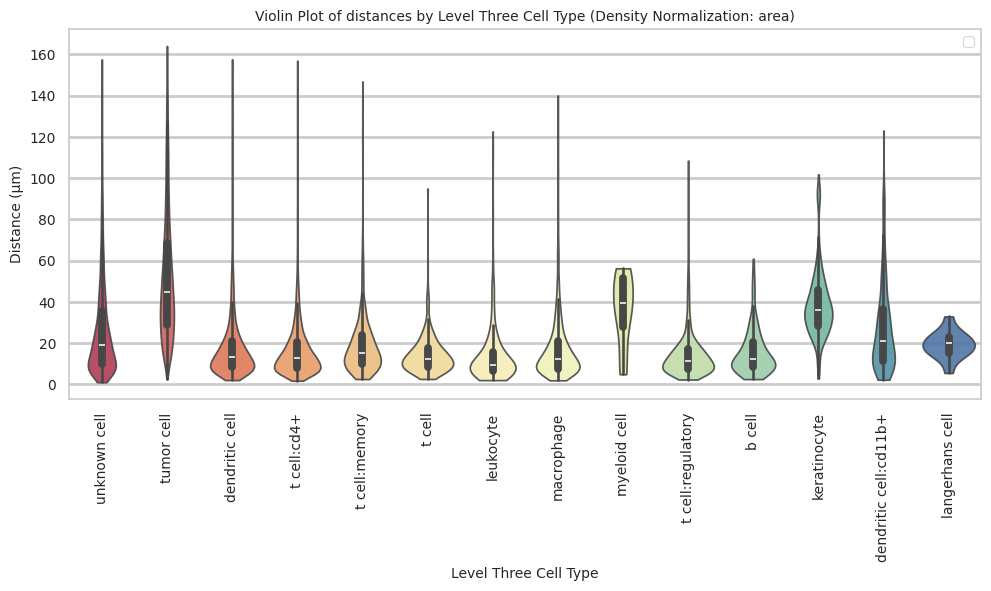
plot_distance_distribution_boxplots_by_region(df_all_edges_with_cell_type_level, cell_type_level, os.path.join(basepath, figures_output_dir))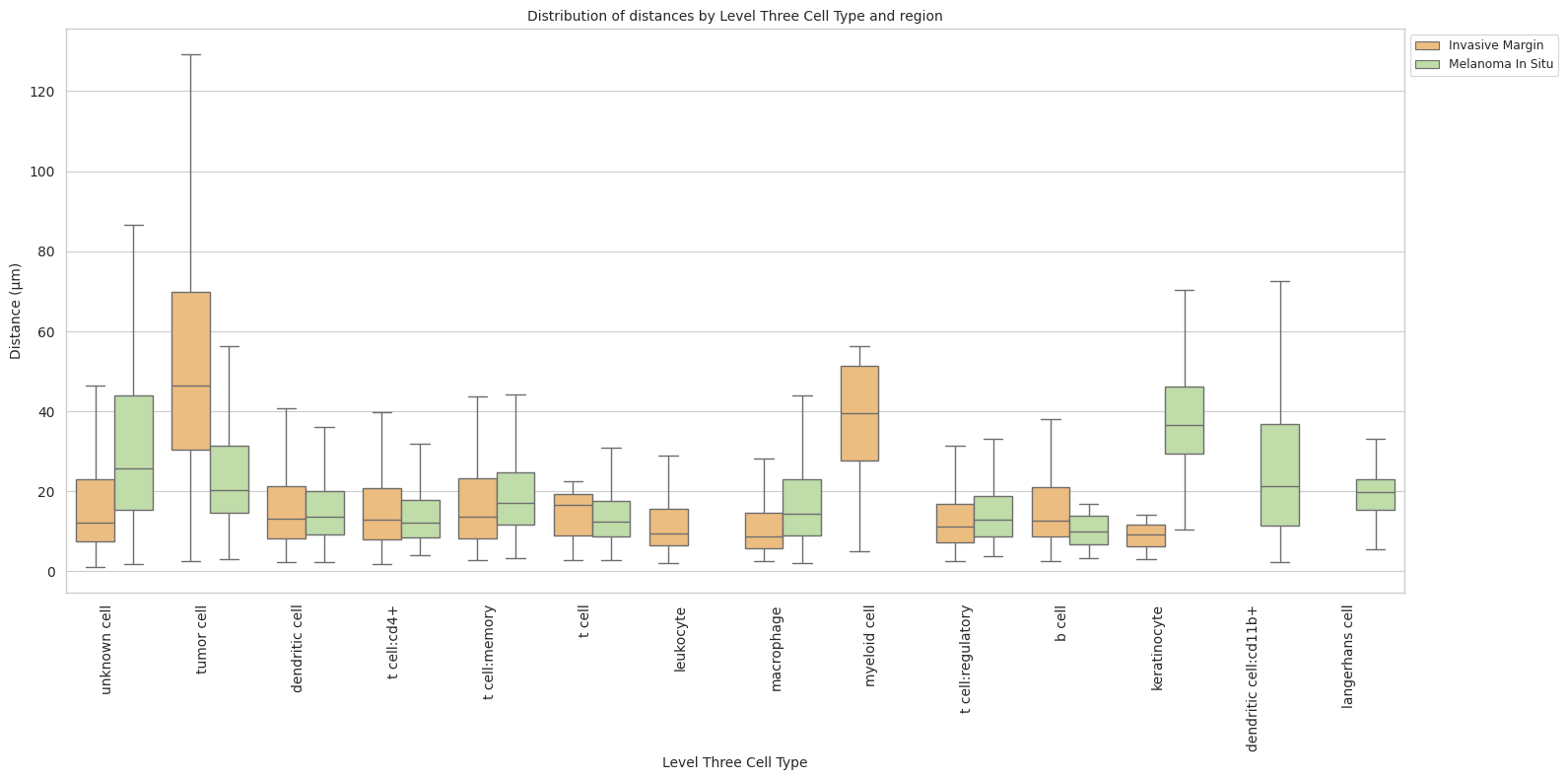
plot_distance_distribution_heatmap(df_all_edges_with_cell_type_level, cell_type_level, os.path.join(basepath, figures_output_dir))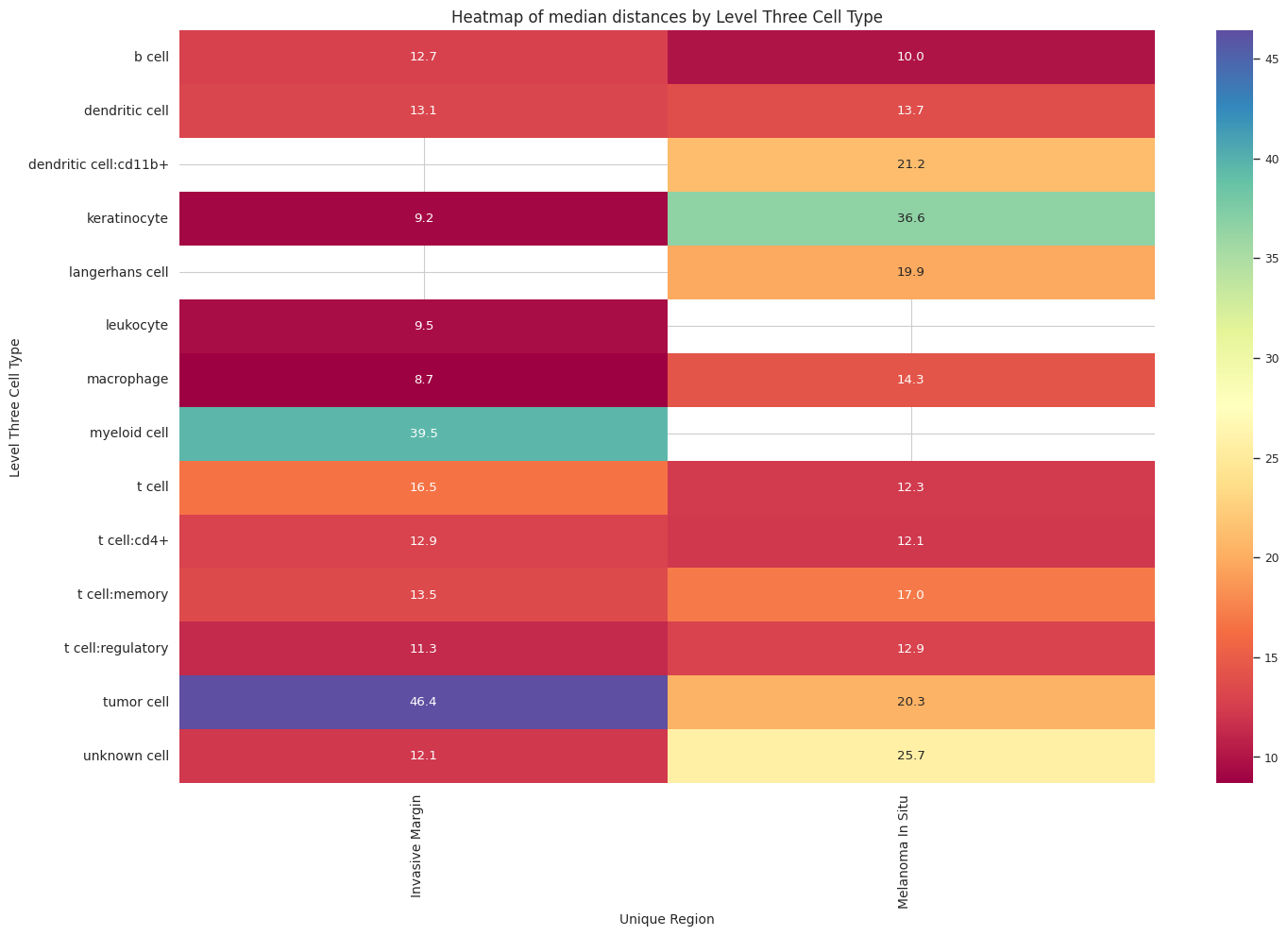
plot_violin_plots_all_regions(df_all_edges_with_cell_type_level, cell_type_level, os.path.join(basepath, figures_output_dir), density_norm="count") # Or, density_norm="count" or "area" based on preference.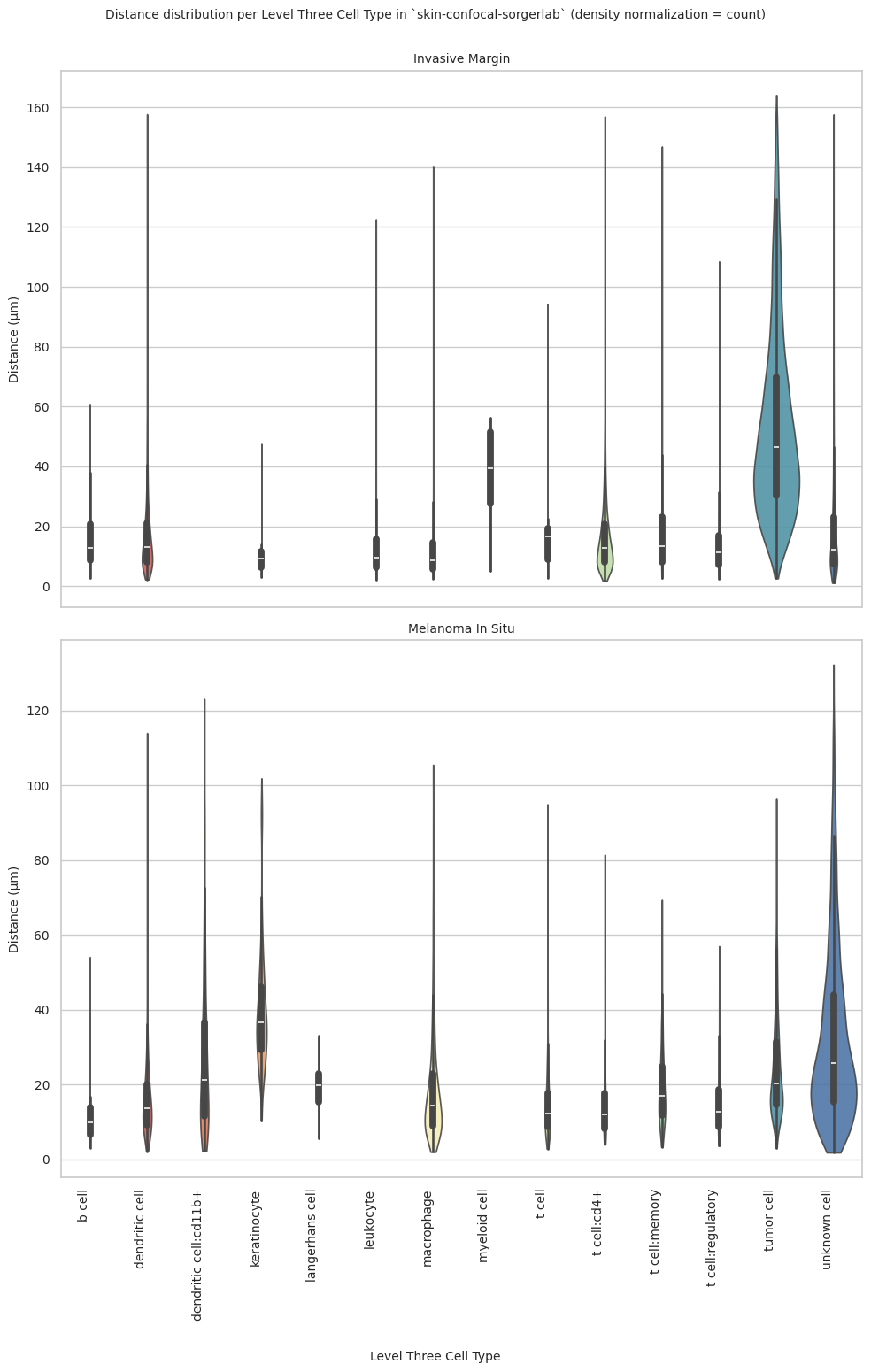
plot_violin_cells_per_celltype_split_by_condition(df_all_edges_with_cell_type_level, cell_type_level, os.path.join(basepath, figures_output_dir), density_norm="count") # density_norm="count" or "area" can be used based on preference.